A Role For BATF3 in Tu9 Differentiation And T-cell-driven L Pathologies MuCOsal(Part 1)
Jun 13, 2022
To learn more info plz contact david.wan@wecistanche.com
T helper 9 (T,9) cells are important for the development of inflammatory and allergic diseases. The TH9 transcriptional network converges signals from cytokines and antigen presentation but is incompletely understood. Here, we identified TL1A, a member of the TNF superfamily, as a strong inducer of mouse and human TH9 differentiation. Mechanistically, TL1A induced the expression of the transcription factors BATF and BATF3 and facilitated their binding to the //9 promoter leading to enhanced secretion of I-9, BATF-and BATF3-deficiencies impaired IL-9 secretion under Tμ9 and Ty9-TL1A-polarizing conditions. In vivo, using a T-cell transfer model, we demonstrated that TL1A promoted IL-9-dependent, T,9 cell-induced intestinal and lung inflammation. Neutralizing IL-9 antibodies attenuated TL1A-driven mucosal inflammation. Batf3-7T_9-TL1A cells induced reduced inflammation and cytokine expression in vivo compared to WT cells. Our results demonstrate that TL1A promotes T_9 cell differentiation and function and defines a role for BATF3 in T-cell-driven mucosal inflammation.

Click here to learn more about Cistanche
INTRODUCTION
Specialized subsets of T helper (T-) cells play an integral role in maintaining tissue homeostasis and during the development of inflammatory diseases at mucosal surfaces. TH9 cells have been recently identified as an independent T cell subset that produces mainly IL-9 but also IL-10 and IL-21.12 Tμ9 cells have been implicated in several diseases, including allergic lung inflammation, experimental autoimmune encephalomyelitis (EAE), colitis, 3- Recently, TH9 cells have had parasitic worm infections and cancer. been reported to play a role in inflammatory bowel diseases (IBD).-9Ulcerative colitis (UC) patients have elevated numbers of mucosal IL-9+ T cells and the IL-9 receptor (IL-9R)is upregulated on the intestinal epithelium. IL-9 deficiency suppresses the development of acute and chronic oxazolone-induced colitis, a model that mimics UC. In Crohn's disease (CD) patients, high-8-10 serum IL-9 levels correlate with severe disease.
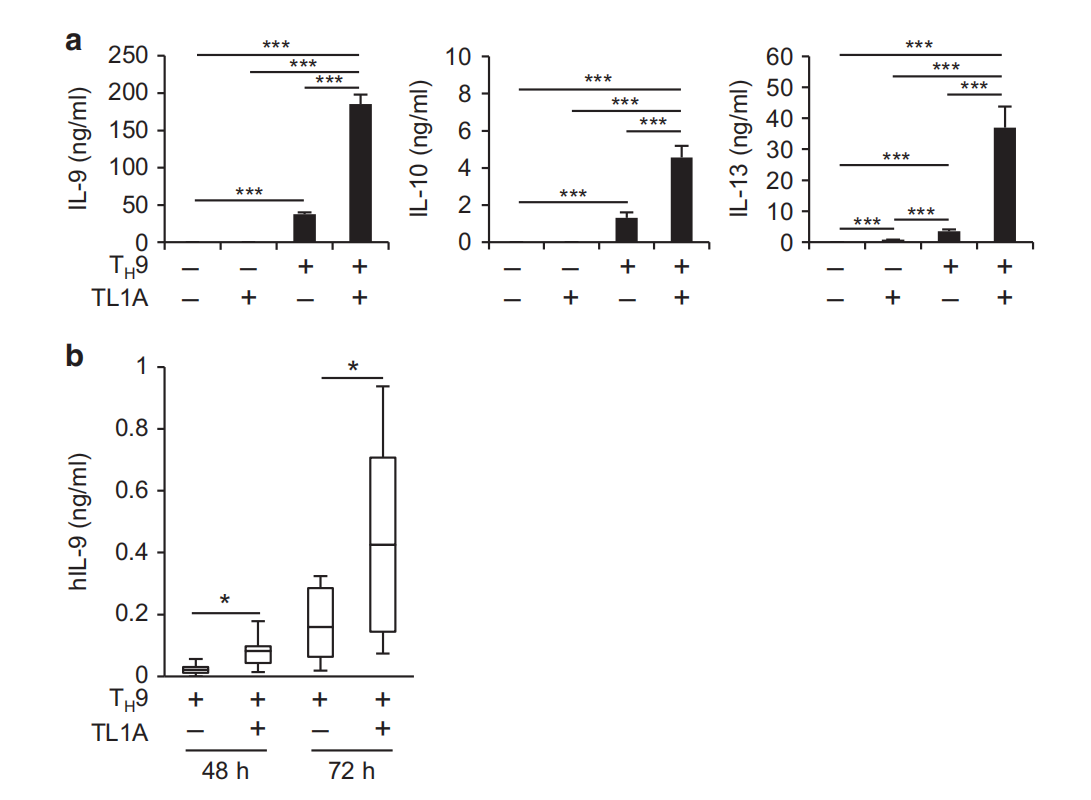

Fig.1 TL1A enhances the differentiation of murine and human TH9 cells. Naive murine CD4+ T cells were differentiated under Ty0-or T9-polarizing conditions with or without TL1A for 3 days, and an ELJSA analysis of ]L-9, IL-10, and IL-13 secretion. b Naive human CD4 T cells (CD4'CD45RACD45ROCD25) were isolated from PBMCs of healthy donors and differentiated under TH9-polarizing conditions with or without TL1A for 3 days. ELISA analysis of human IL-9 secretion at 48 and 72 h. Box-whisker plot showing the IL-9 concentrations in culture supernatants for TH9 and TH9-TL1A-treated cells. The boxes represent the median and 25th and 75th percentile, and the whiskers represent the minimum and maximum values of the distribution. N=7-9. Data represent means± SD of one independent experiment out of four (independent experiments. *p< 0.05,***p <0.005 determined by Student's t-test
TH9 cells differentiate from naive CD4+T cells in the presence of TGF-β1 and IL-4. Several transcription factors downstream of the T-cell receptor(TCR), TGF-β1, and IL-4 receptors are required for the differentiation of TH9 cells including IRF4, STAT6, GATA3, PU.1, NF-kB, and basic leucine zipper transcription factor ATF-like (BATF.11-15 However, the transcriptional program and inflammatory triggers that drive the differentiation of Tμ9 cells are still not well understood and a lineage-defining factor associated with lL-9 expression has not been identified.
The family of BATF transcription factors is comprised of BATF, BATF2, and BATF3 and belongs to the AP-1 family of transcription factors that include JUN and FOS. Transcription factors in the BATF family are composed of a DNA-binding domain and leucine zipper motif but lack a transactivation domain and were initially described as negative regulators of AP-1 activity." However, recent studies have demonstrated that BATF family members interact with IRF transcription factors including IRF4 and IRF8, and bind to AP-1-IRF composite element sequences to regulate their target genes in T cells and dendritic cells. BATF has been shown to be required for the development of Ty9 cells as well as TH2, TA17, and Try cells.15,19BATF3 has been described to control the development of CD8a+ and CD103+ dendritic cells. "2' In contrast to BATF, a nonredundant function of BATF3 in the development of Tu cells has not been demonstrated. In addition, the extracellular stimuli that can activate the BATF3 pathway in CD4+ Tu cells remain to be elucidated.
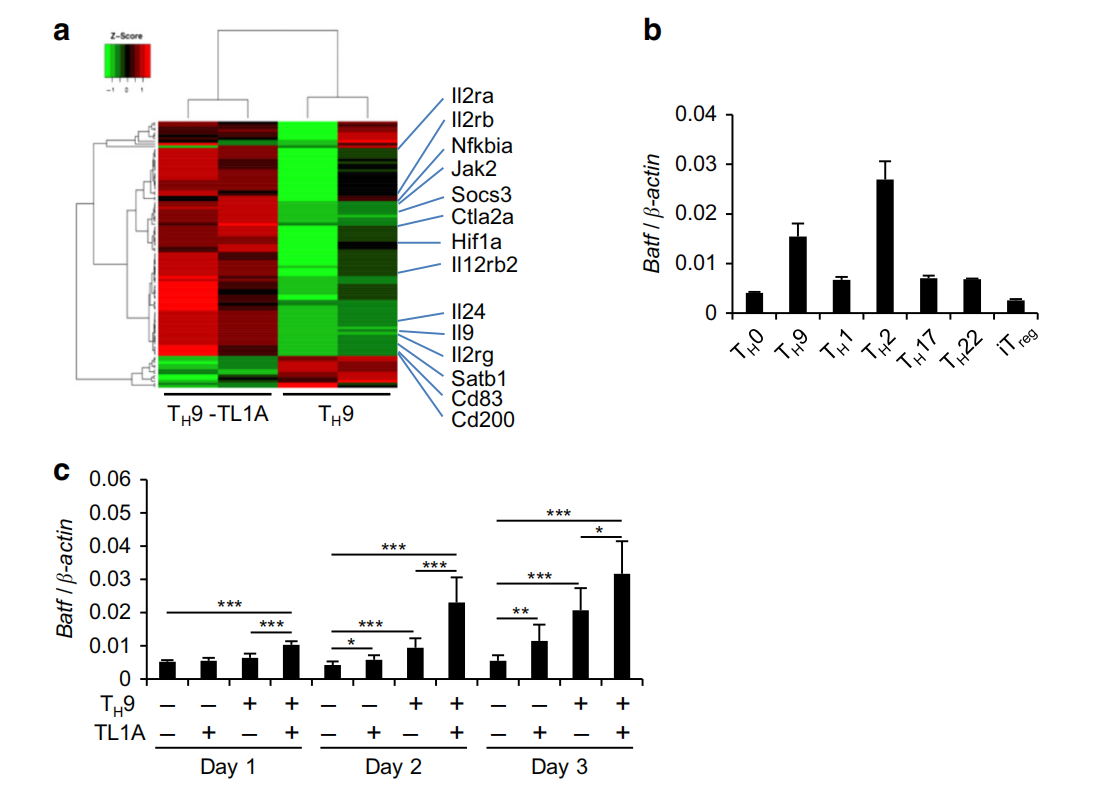
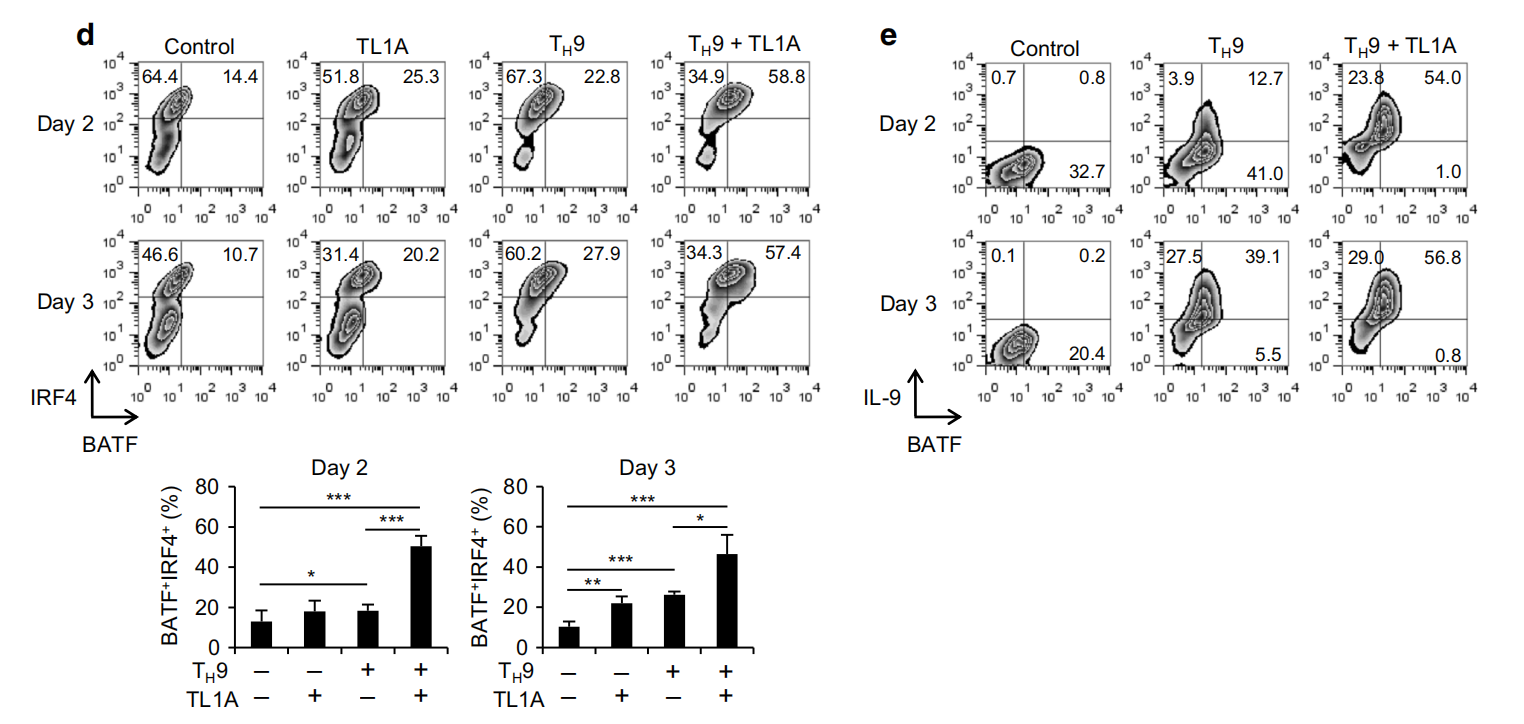
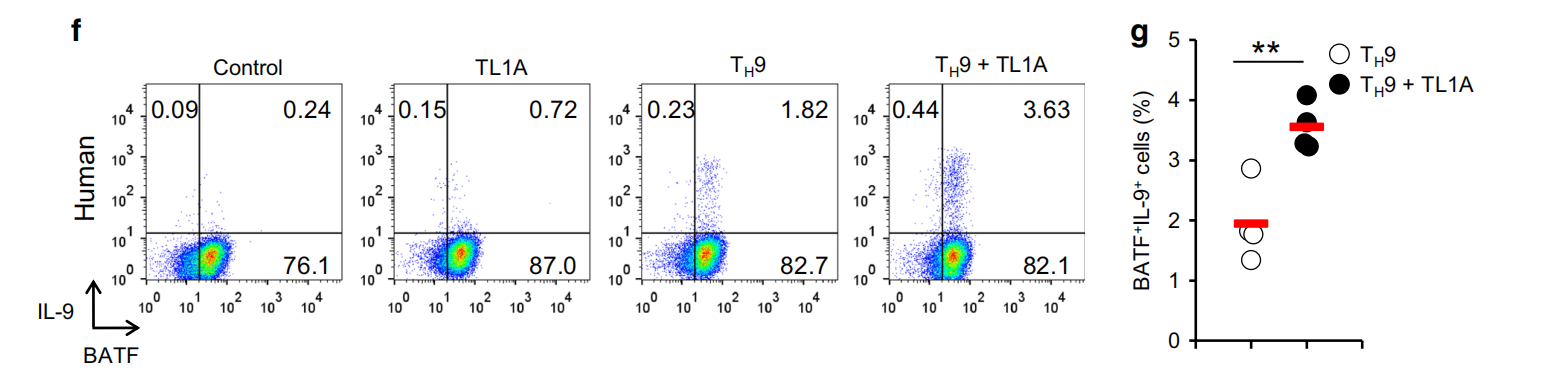
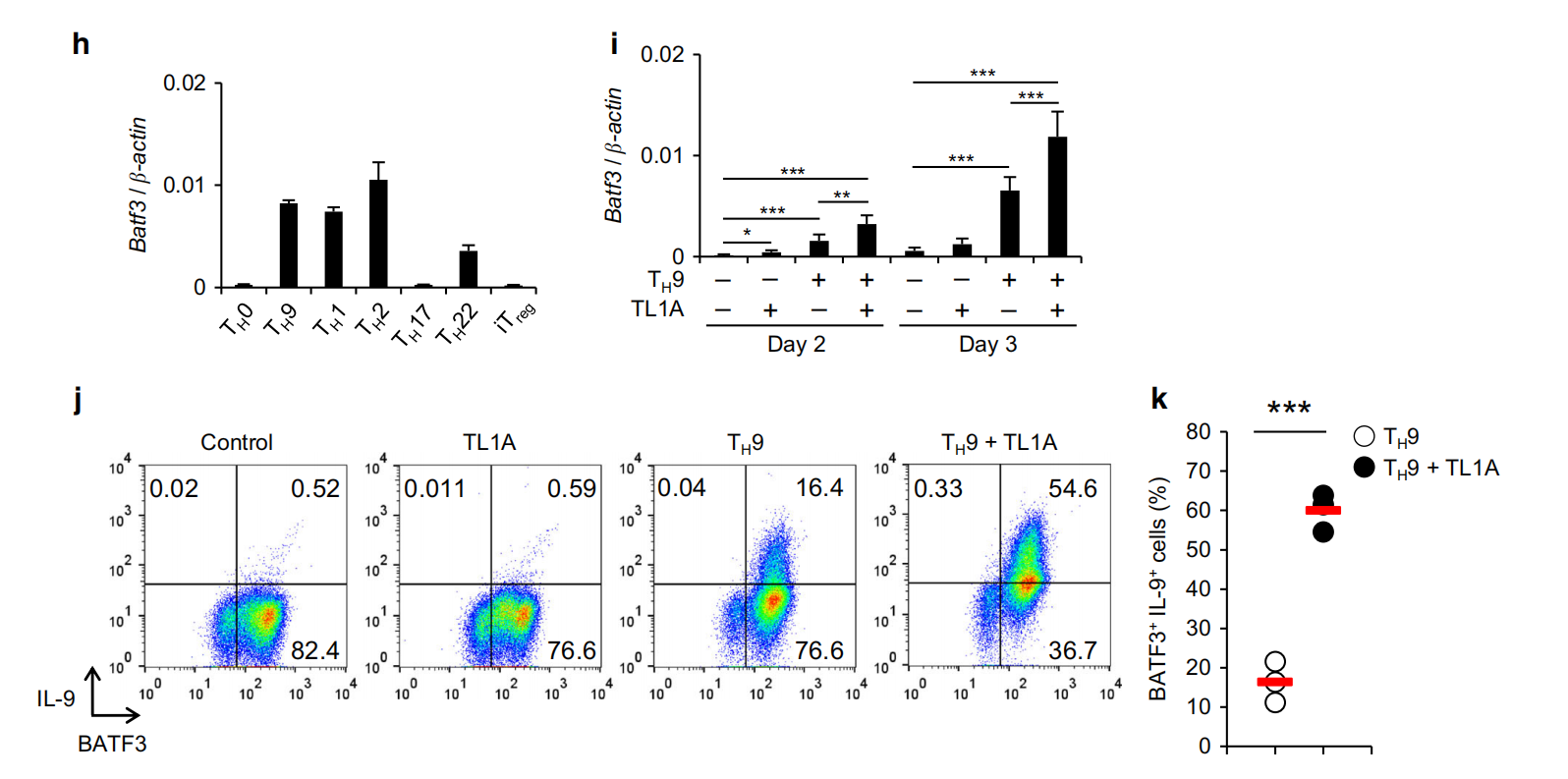
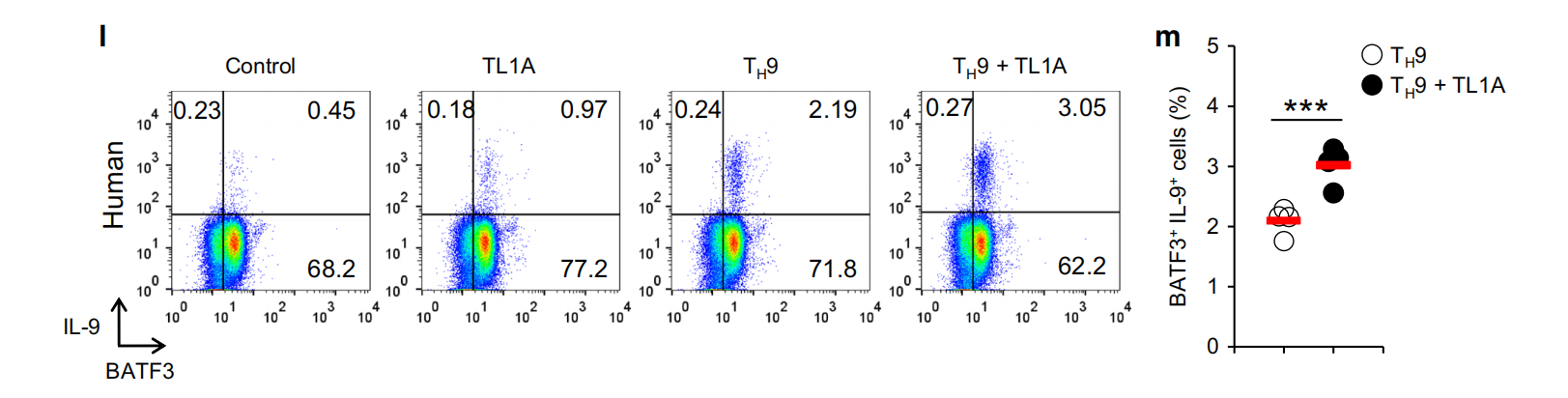
Fig.2 TL1A upregulates BATF and BATF3 expression during T,9 differentiation. Transcriptional profiling of TH9 and TH9-TL1A cells by RNA sequencing. Heat map displaying RNA-sequencing data using the top 100 genes with the largest IQR (interquartile range). The dendrograms to the left and above the heat map represent the clustering of genes (rows) and samples (columns). b Naive CD4+ T cells were differentiated into different TH subsets and Tregs for 3 days. Relative BATF mRNA expression was analyzed by PCR,c Naive CD4+T cells were differentiated under TH0- or TH9-polarizing conditions with or without TL1A for the indicated time periods. PCR analysis of relative BATF mRNA expression,d Co-staining of BATF and IRF4(top). Quantitative analysis of the percentages of BATFIRF4 cells(bottom).e Co-staining of BATF and IL-9. f-g
TL1A. f Representative intracellular staining of IL-9 and BATF at 48h.g Frequency of BATFfIL-9+ T cells differentiated under Ty9 and T9-TL1A conditions. Means are shown. Each symbol represents an individual donor. N=4. h Relative Batf3 mRNA expression in different murine Th subsets and Tregs was analyzed by qPCR.I Relative Batf3 mRNA expression at indicated time points. j Representative co-staining of BATF3 and IL-9. k Frequency of BATF3IL-9+ T cells differentiated under Ty9 and TH9-TL1A conditions. Means are shown. Each symbol represents an individual experiment. N=3. I-m Naive human CD4+ T cells were differentiated under T,9-polarizing conditions with or without TL1A. I Representative intracellular staining of IL-9 and BATF3 at 48 h. m Frequency of BATF3L-9+ T cells differentiated under TH9 and TH9-TL1A conditions. Means are shown. Each symbol represents an individual donor. N=4. Data represent means±SD of one independent experiment ut of two(a) or at least three (b-d. i. k) independent experiments.*n <0.05. n <0.01.***n < 0.005 as determined by Student's t-test
We have recently shown that TL1A, a TNF superfamily member that plays an important role in immune-mediated diseases including IBD, 25 induces the secretion of IL-9 in Tu17 cells. "Furthermore, a recent publication demonstrated that TL1A, via its receptor DR3, promotes TA9 differentiation through IL-2- and STAT5-dependent but PU.1- and STAT6-independent mechanisms in allergic lung inflammation.2’ However, the transcriptional programs induced by TL1A and the pathogenicity of TL1A-induced TH9 cells in intestinal inflammation and IBD remains to be elucidated.
Here, we demonstrate that TL1A promotes the differentiation of human and murine TH9 cells via a novel signaling pathway. We define BATF3 as a novel transcriptional regulator for the differentiation of Tp9 cells. TL1A leads to chromatin remodeling at the lI9 locus and recruitment of the pioneer transcription factors IRF4, and BATF, which are important components for IL-9 transcriptional activation, and BATF3 to conserved regions within the 9 promoters. Furthermore, TL1A upregulates the expression of BATF and BATF3 in a STAT6-dependent manner.BATF- and Batf3-deficient T-9 and TL1A-induced TH9(T-9-TL1A) cells are impaired in their IL-9 production. TH9-TL1A cells are highly pro-inflammatory in vivo when adoptively transferred into Ragi-7-mice as evident by severe mucosal inflammation in the intestines and lungs, which was attenuated by treatment with neutralizing IL-9 antibodies. Adoptive transfer of Batf3-/TH9-TL1A cells into Rag1-7mice leads to significantly reduced mucosal inflammation and cytokine production in comparison with WT TH9-TL1A cells. Our data demonstrate a novel role for TL1A and BATF3 in developing pathogenic effector T,9 cells and identify this signaling pathway as a potential therapeutic target in T_9-driven pathologies, including IBD and allergic lung disease.

RESULTS
TL1A enhances murine and human Tμ9 cell differentiation in vitro To determine the effect of TL1A on TH9 differentiation, we stimulated naive CD4 T cells under TH9 conditions with or without TL1A. TL1A significantly enhanced IL-9 secretion and I/9 mRNA expression (Fig. 1a, Supplementary Figure 1a). TL1A alone did not induce IL-9 secretion, suggesting that TL1A acts synergistically with TGF-β1 and IL-4 to promote IL-9 production. Secretion and mRNA expression of T-9-associated cytokines L-10 and IL-13 were also enhanced by TL1A (Fig.1a, Supplementary Figure 1a, b). Consistent with previous reports, time-course experiments of intracellular IL-9 staining revealed transient IL-9 expression with a maximal induction under T-9 conditions on day 3 and a decline back to baseline on day 5(Supplementary Figure 1b).28,2TL1A induced a significantly higher percentage of IL-9-producing cells throughout the time course of the experiments with a shift in kinetics toward an earlier and stronger IL-9 induction that even at an early time point exceeded the maximum of IL-9 production induced under TH9 conditions (Supplementary Figure 1b). We also observed an increase of IL-9+IL-10+ cells in the presence of TL1A (Supplementary Figure 1b). We did not observe any shift toward other Tu subsets (Supplementary Figure 1c). The enhancement of TH9 differentiation by TL1A was completely dependent on its receptor DR3 (Supplementary Figure 1d). Taken together, these data suggest that TL1A synergizes with TGF-β1 and IL-4 in TA9 differentiation and IL-9 production.
TL1A also significantly enhanced TH9 differentiation in an antigen-specific setting using OVA-specific OT-ll cells (Supplementary Figure 1e, f). TL1A did not affect cell proliferation during TH9 differentiation (Supplementary Figure 1g), as has been demonstrated for memory CD4+ T cells. " These data suggest that TL1A enhanced TH9 differentiation without affecting proliferation. Furthermore, we examined IL-9 production in cells that were differentiated under Ty9 and TH9-TL1A conditions for 3 days and restimulated with anti-CD3s/CD28. We observed increased IL-9 production in TH9-TL1A cells compared to TH9 cells after secondary stimulation that was further enhanced in the presence of TL1A (Supplementary Figure 1h, i). These data suggest that TL1A also enhanced IL-9 production from differentiated TH9 cells. Next, we determined the effect of TL1A on the differentiation of human Tu9 cells. Naive CD4+ T cells were isolated from PBMC from healthy donors and stimulated under Tu9 condition in the presence of human TL1A. Consistent with our data from mouse TH9 cells, the addition of TL1A significantly enhanced IL-9 secretion from human TH9 cell cultures (Fig. 1b).
TL1A induces the expression of the transcription factors BATF and BATF3 To elucidate signaling pathways induced by TL1A, we used RNA sequencing to assess the transcriptomic profile in TH9 and TH9-TL1A cells. TL1A did not substantially alter the TH9 transcriptional profile indicating that TL1A did not globally skew TH9 differentiation (data not shown). However, 219 genes were differentially expressed in TH9-TL1A cells(Fig. 2a, Supplementary Tables 1 and 2). Pathway enrichment analysis identified significant changes in genes involved in cytokine-cytokine receptor and JAK-STAT signaling pathways in T,9-TL1A conditions(Supplementary Tables 1 and 2, data not shown). BATF3 and several BATF-regulated genes were significantly upregulated under T,9-TL1A conditions (Supplementary Table 3) suggesting that the transcription factors BATF3 and BATF might be involved in the differentiation of TH9-TL1A cells. BATF has been described to be required To determine the development of Tp17 and TA9 cells.15,31-34 -whether TL1A activates BATF family members, we first assessed the expression level of BATF in different Tu subsets and detected strong BATF mRNA expression in TH9 and TH2 cells (Fig. 2b). TL1A further enhanced the expression of BATF under Tu9 conditions, particularly on day 1 when TH9 conditions alone are not sufficient to induce BATF mRNA (Fig.2c). TL1A alone induced BATF mRNA to a similar degree as Tp9 conditions (Fig. 2c). Intracellular staining for BATF confirmed that Ty9-TL1A stimulation significantly increased the percentage of BATF+ cells (Fig. 2d). Furthermore, TL1A alone increased the percentage of BATF+ cells and more importantly the percentage of BATF*IRF4+ cells compared to TH9 conditions suggesting that TL1A might enhance the formation of IRF4-BATF cooperative complexes (Fig. 2d). TH9-TL1A stimulation also enhanced the percentage of BATF+IL-9+ cells (Fig. 2e).
In addition, human TH9 cells had a significantly higher percentage of IL-9-producing cells when stimulated with TL1A (Fig. 2f, g). Notably, the majority of IL-9-producing cells co-expressed BATF. Co-staining of BATF and IRF4 showed that the addition of TL1A increased the percentage of BATF*IRF4+ cells during differentiation of human T-9 (data not shown), suggesting that the enhancement of IL-9 production via the BATF signaling pathway by TL1A is conserved between mouse and human TH9 cells. In contrast to BATF, the role of BATF3 in the differentiation of TH cells is less defined. To confirm our RNA-seq data, we determined the expression level of Batf3 in T-9 cells and other TH subsets and found that Batf3 mRNA expression was the highest in T-9, T,1, and TH2 cells (Fig. 2h). TL1A enhanced the expression of Batf3 under T-9 conditions(Fig.2i). Intracellular staining for BATF3 confirmed that T,9-TL1A stimulation significantly increased the percentage of BATF3+ IL-9+ cells (Fig.2j, k).In human T9 cells, TL1A significantly enhanced the expression of BATF3+IL-9+ cells(Fig.2l, m). Similar to BATF, the majority of IL-9-producing cells co-expressed BATF3.
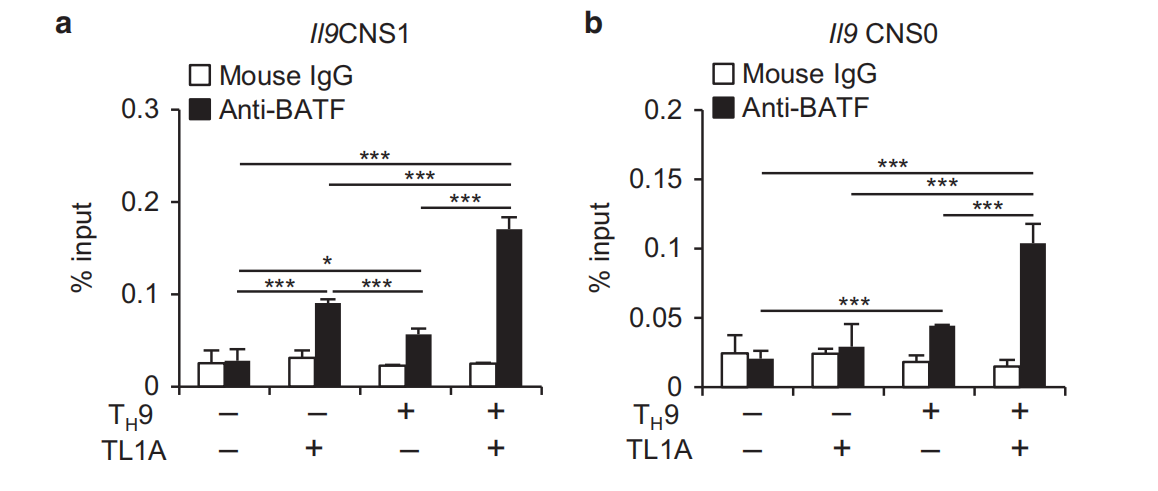
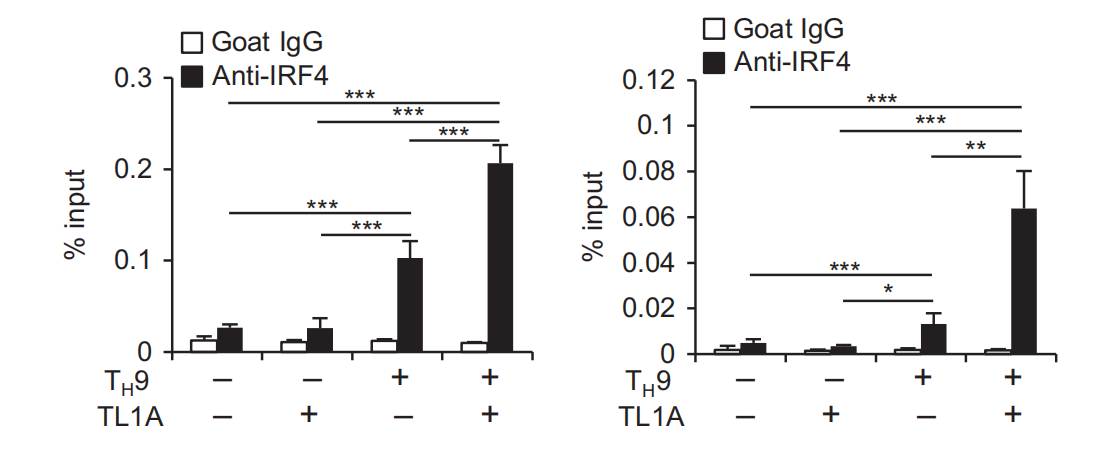

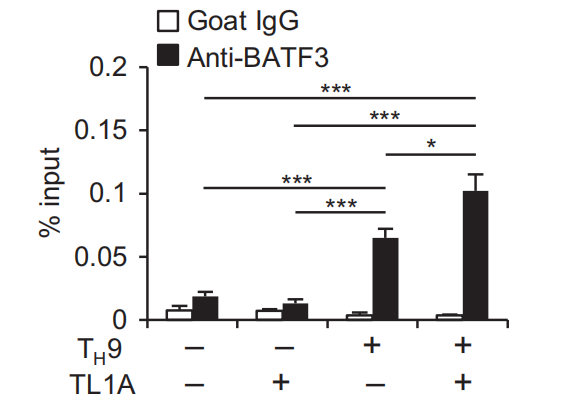

Fig,3 TL1A enhances binding of BATE IRF4, and BATF3 to the //9 promoter during Ty9 differentiation, a, b Naive murine CD4+ T cells were differentiated under TH0- or Tμ9-polarizing conditions with or without TL1A for 2(BATF, IRF4, and AcH3), or 3(BATF3) days. ChP assays were performed for BATF, IRF4, BATF3, or AcH3 binding to conserved noncoding sequences(CNS) 1 (a) and CNS0 (b) of //9. Data represent the means ±SD of one independent experiment out of two independent experiments.*p<0.05,*p< 0.01,**p< 0.005 as determined by Student's t-test
BATF and BATF3 are critical transcription factors in TL1A-induced T_9 differentiation
To further define the role of BATF and BATF3, we performed chromatin immunoprecipitation (ChIP) assays to determine binding at two Il9 conserved noncoding sequences(Il9 CNS1, CNSO).5 TL1A alone induced significantly greater binding of BATF to II9 CNS1 compared to TH9 conditions and TH9-TL1A further enhanced BATF binding (Fig.3a). At the more upstream I/9 CNS0, TL1A also enhanced BATF binding under T,9 conditions while TL1A alone had no effect (Fig.3b). TL1A enhanced IRF4 binding under Ty9 conditions at CNS1 and CNS0(Fig.3a, b). Interestingly, the binding of acetylated histone 3(AcH3)at CNS1 and CNSO that was previously described to facilitate chromatin remodeling at the Il9 locus during TH9 differentiation was also enhanced under Ty9-TL1A conditions (Fig.3a,b). BATF3 binding to CNS1 was upregulated in TH9 cells and the binding was further enhanced under TH9-TL1A conditions while BATF3 did not bind to CNS0 (Fig.3a, data not shown). These data suggest that TL1A enhances BATF, BATF3, and IFR4 binding to the //9 promoter under TH9 conditions to induce ll9 expression. To confirm the roles of BATF and BATF3 in TL1A-dependent and Batf3-7-T cells. IL-9, IL- Ty9 differentiation, we used Batf7- a 10, and IL-13 production was severely impaired in BATF-/-cells under Tμ9 conditions (Fig. 4a-d). However, TL1A stimulation at least partially overcame the requirement for BATF for IL-9 and IL-13 production during Ty9 differentiation (Fig.4a, b, d). In the absence of BATF3, the induction of IL-9 was significantly reduced, particularly under Tμ9-TL1A conditions (Fig.4e,f, Supplementary Figure 2a).In contrast to BATF, BATF3 was not required for-10 or IL-13 expression (Fig.4g, h, Supplementary Figure 2b, c).To define the signaling pathways that lead to BATF and BATF3 upregulation in the presence of TL1A, we used Stat6-7and p50-/cells. p50-/TH9 cells had reduced percentages of BATF+IRF4t cells and the increase in the percentages of BATF+IRF4+ cells under TL1A stimulation was almost completely abolished (Supplementary Figure 3a, b). Upregulation of BATF and Batf3 mRNA under T.9 conditions was p50-independent, while the enhancement of BATF and Batf3 expression under TH9-TL1A conditions was completely p50-dependent (Supplementary Figure 3c, d). Stat6-/T9 cells had reduced percentages of BATFIRF4+cells and their increase under Tμ9-TL1A stimulation was completely abolished(Supplementary Figure 4a, b). Similarly, upregulation of BATF and Batf3 mRNA was completely dependent on STAT6 under TH9 and TH9-TL1A conditions (Supplementary Figure 4c, d). We observed no significant effects of BATF or BATF3 deficiency on the expression of Batf3 and BATF, respectively, suggesting that there is no compensation between the two transcription factors (Supplementary Figure 5). Together, these data suggest that BATF and BATF3 play important and distinct roles during T_9 differentiation in the presence of TL1A and that increased BATF and BATF3 expression under TL1A stimulation is dependent on STAT6 and the canonical NF-KB pathway. Next, we performed RNA sequencing on WT and Batf3-/-Tμ9-TL1A cells. Totally,139 genes were differentially expressed in WT versus Batf3-7Ty9-TL1A cells(Fig.4i, Supplementary Table 4). Gene ontology pathway enrichment analysis identified significant changes in genes involved in the regulation of transcription, cell proliferation, and intracellular signal transduction (Supplementary Tables 5 and 6). These results reveal a role for BATF3 in the differentiation of Ty9-TL1A cells. T-9-TL1A cells are highly pathogenic in vivo To determine the pathogenic potential of TH9-TL1A cells in vivo, we adoptively transferred ex vivo-differentiated CD45.1+Tμ9 or TA9-TL1A cells into Rag1-'-mice. Mice that received Ty9 cells did not develop significant weight loss (Fig. 5a). However, mice that received TH9-TL1A cells lost weight and developed more severe intestinal inflammation, particularly in the small intestine (Fig. 5a-c). We also observed severe pulmonary inflammation, characterized by peribronchial and perivascular cellular infiltration of inflammatory cells in TA9-TL1A recipients. Staining with Alcian Blue periodic acid-Schiff (AB-PAS) showed hyperproliferation of mucin-producing cells in the airway epithelium of Ty9-TL1A recipients (Supplementary Figure 6a, b). We observed higher cellularity in spleens and lamina propria mononuclear cells (LPMC), and higher percentages and absolute numbers of splenic, MLN, and LPMC CD4+ cells in TH9-TL1A recipients (Fig. 5d, e)suggesting that Ty9-TL1A cells have a higher proliferative potential in vivo. Ki67 staining confirmed that cells from MLN, spleen and LPMC from TH9-TL1A recipients are more proliferative even 6 weeks after T-cell transfers (Fig. 5f). Significantly higher secretion of IL-13 and IL-17 in anti-CD3/anti-CD28-restimulated MLN cells from TH9-TL1A recipients was observed (Fig. 5g), and significantly higher secretion of IL-9 and IL-13 in anti-CD3/anti-CD28-restimulated LPMC from Ty9-TL1A recipients (Fig. 5h). We observed a higher absolute number of splenic and MLN IL-9+, IL-13+, and IL-17+ cells and a higher absolute number of MLN, splenic, and LPMC CCR6+ and CD103+ T cells in TH9-TL1A recipients consistent with a role of these cells at mucosal surfaces (Fig.5i,j). IL-9 is required for in vivo inflammation driven by T#9-TL1A cells To examine whether increased IL-9 production by TH9-TL1A cells contributed to intestinal and lung inflammation, we treated mice receiving TH9-TL1A cells with neutralizing IL-9 or control antibodies for the duration of the T-cell transfer experiment. Histopathological analysis demonstrated significantly reduced intestinal inflammation in anti-lL-9-treated mice compared to isotype controls (Fig. 6a, b). We also observed attenuated lung inflammation with decreased peribronchial infiltrates and a reduction in mucus-producing cells in anti-lL-9-treated mice (Supplementary Figure 6c). Total cell numbers and total CD4tCD45.1+T cells in MLN were reduced in anti-IL-9-treated mice compared to isotype controls (Fig. 6c). Anti-IL-9 treatment also reduced the production of IL-13 and IL-17 in MLN and spleens, while IFN-y production was unchanged (Fig. 6d, data not shown). Together, these data demonstrate that T-9-TL1A cells were potent effector cells and resulted in intestinal and lung inflammation in vivo and that the pathogenic effect was IL-9-dependent.
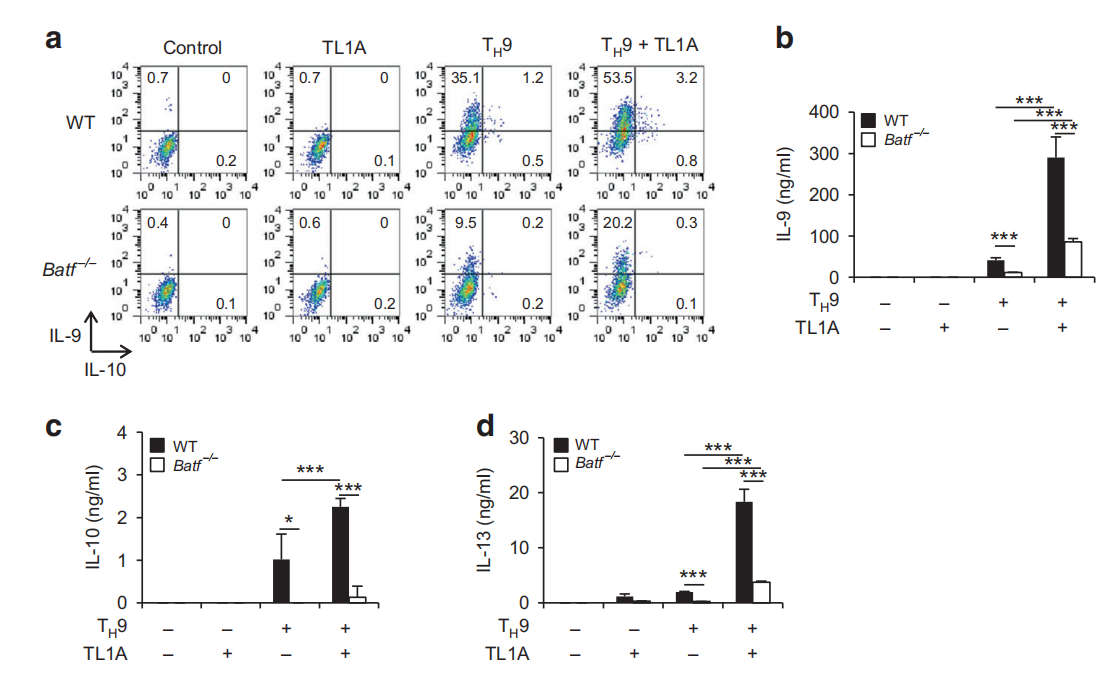
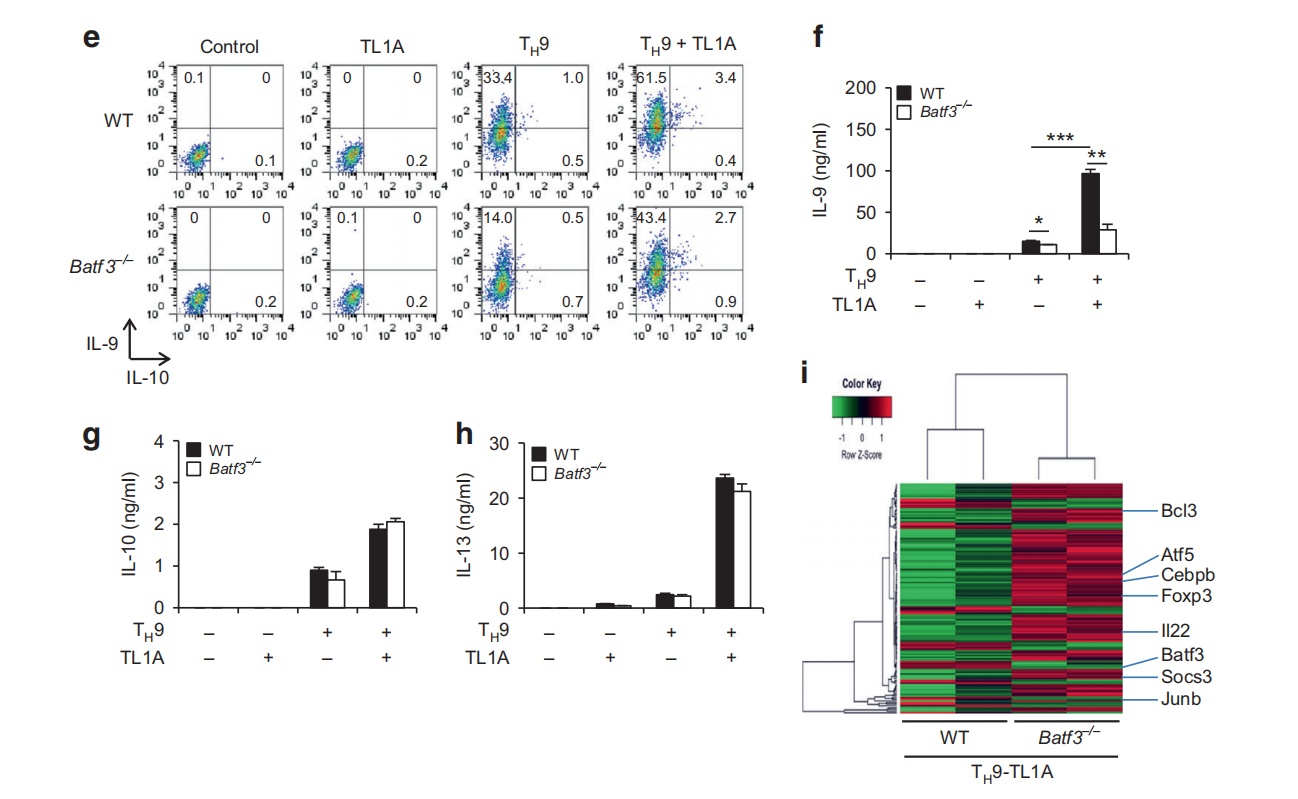
Fig.4 Effects of BATF and BATF3 deficiencies on TH9 differentiation. a-d Naive CD4 T cells from Batf7or WT mice were differentiated under TH0- or TH9-polarizing conditions with or without TL1A for 3 days. Intracellular staining of IL-9 and IL-10.b ELISA analysis of IL-9 production.c ELISA analysis of IL-10 production.d ELISA analysis of IL-13 production. e-h Naive CD4+ T cells from Batf3-7-or WT mice were differentiated under T,9-polarizing conditions with or without TL1A for 3 days. e Intracellular staining of IL-9 and IL-10.f ELISA analysis of IL-9 production. g ELSA analysis of IL-10 production. h ELISA analysis of IL-13 production. I Transcriptional profiling of WT and Batf3′TH9-TL1A cells by RNA sequencing. Heat map displaying RNA-sequencing data of genes differentially expressed with p<0.01. The dendrograms to the left and above the heat map represent the clustering of genes (rows) and samples (columns). Data represent means± SD of one independent experiment out of two (i) or three (a-h) independent experiments.*p<0.05,**p < 0.01, ***p <0.005 determined by Student's t-test
BATF3 contributes to in vivo inflammation driven by TH9-TL1A cells To determine whether BATF3 plays a role in Tu9-driven pathologies in vivo, we adoptively transferred ex vivo-differentiated WT or Batf3-/-TA9-TL1A into Rag1-/-mice. Mice that received Batf3-'- TA9-TL1A cells develop significantly less pulmonary (Fig. 7a, b, Supplementary Figure 6e) and a trend toward reduced intestinal inflammation (Histoscore: 3.6 vs. 1.2)(Fig. 7a). We observed reduced cellularity in lungs and LPMC, and lower percentages of MLN and lung CD4+ T cells in Batf37′-TL1A recipients compared to WT TH9-TL1A recipients (Fig. 7c, Supplementary Figure 6d). Significantly, lower percentages of IL-13+and IL-17+ cells in spleens, MLN, and lungs from Batf3-'-Ty9-TL1A recipients were observed (Fig. 7d). Collectively, our data indicate that BATF3 plays a role in IL-9 production and T#9 development in vitro and in Tp9-TL1A-driven mucosal inflammation in vivo.

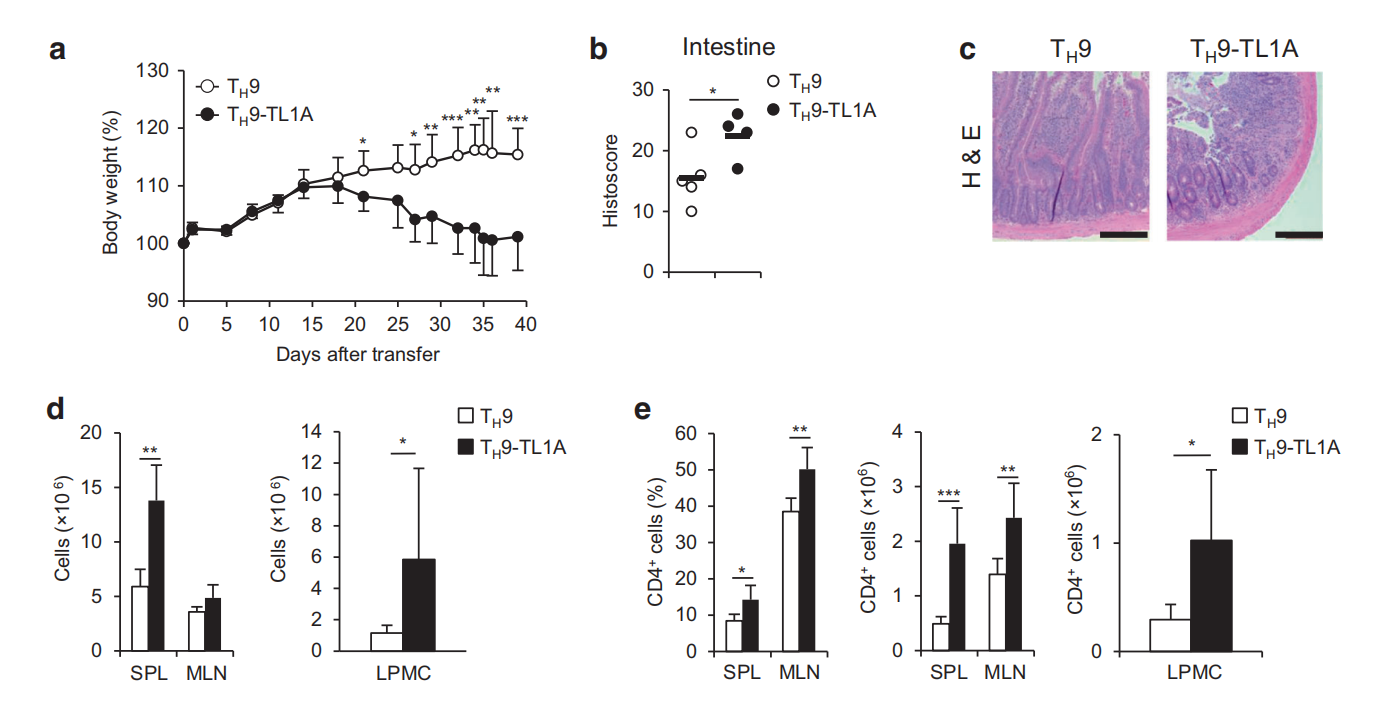

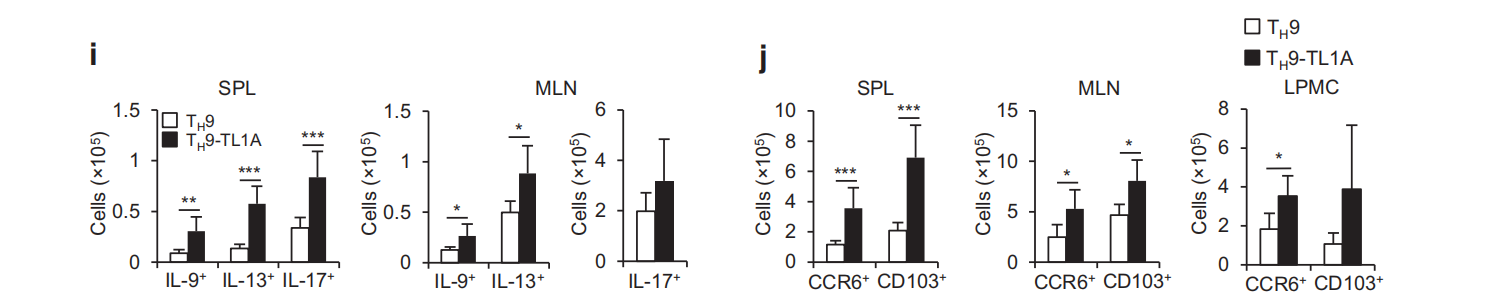
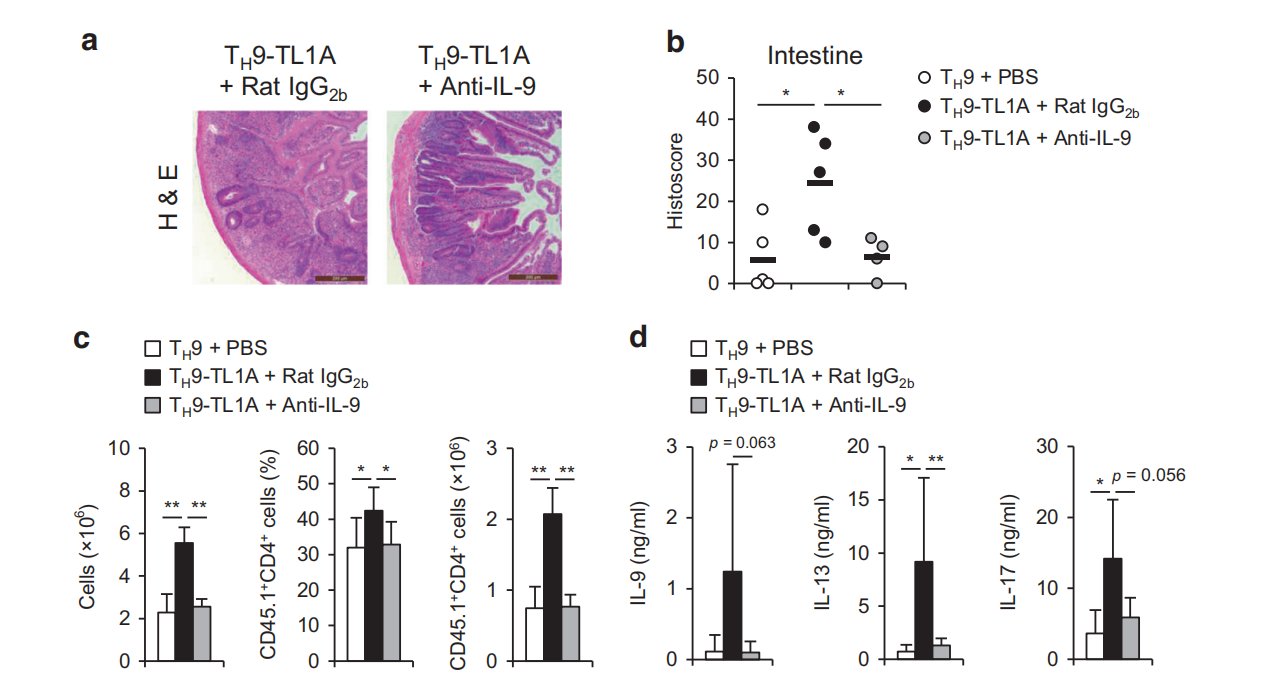
Fig.6 IL-9 is required for in vivo mucosal inflammation driven by Ty9-TL1A cells. T,9 or Ty9-TL1A cells were injected into Rgg1-7mice, and mice were treated three times per week with anti-IL-9 antibody or isotype control, a Representative H&E stainings of the duodenum, Scale bar;200 μm. b Histology scores for the intestine.c Total MLN cell counts (left), frequency of CD45.1+ cells (middle), and total cell counts of CD45.1+cells (right).d ELISA analysis of IL-9, IL-17, and IL-13 production from MLN after ex vivo restimulation with anti-CD3 and anti-CD28. Data represent means± SD of one independent experiment out of two independent experiments. n=4-5/group.*p<0.05,*p<0.01, as determined by Student's t-test (b) or Mann-Whitney U test (c, d)





