Part Ⅱ The Usage Of Antibiotics By COVID-19 Patients With Comorbidities: The Risk Of Increased Antimicrobial Resistance
May 10, 2023
Results
1. Distribution of Clinical Bacterial Isolates among Coronavirus Disease 2019 (COVID-19) Patients
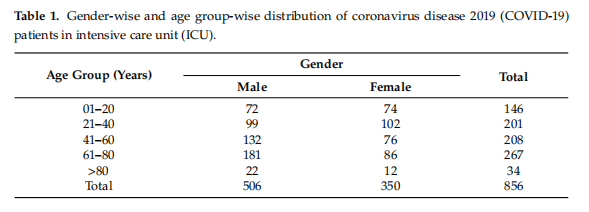
From the total 856 patient samples, n = 506 (59.11%) were male and n = 350 (40.88%) were females as shown in Table 1. A total of 342 (39.95%) samples were found to be positive for different bacterial isolates. Most of the bacterial cultures were found to be positive in urine (n = 136, 39.76%) and tracheal aspirate cultures (n = 51, 14.91%) followed by broncho-alveolar lavage (n = 48, 14.03%), wound swab (n = 37, 10.81%), blood (n = 32, 9.35%), sputum (n = 24, 7.01%), pus swab (n = 5, 1.46%), Foley’s catheter tip (n = 4, 1.16%), abscess (n = 3, 0.87%), and pleural fluid (n = 2, 0.58%).
2. Clinical Isolates from Various Infections
The most common diagnosed comorbidity was chronic kidney disease (CKD), along with urinary tract infection (UTI) (21.4%), which was caused mainly by E. coli (50%), followed by the Klebsiella pneumoniae (20%). The second most diagnosed comorbidity was sepsis and hematuria (19.1%), also caused mainly by E. coli (52.9%), followed by Klebsiella pneumoniae (11.8%) and Proteus spp. (11.7%), respectively. The least common diagnosis was respiratory tract infections (1.11%), caused mainly by Klebsiella pneumoniae (100%). The BSI followed by the lung abscesses (1.07%) was caused mainly by the Klebsiella pneumoniae. (100%). The prevalence of bacterial isolates is shown in Table 2. Supplementary Materials: Figure S1 shows the beta hemolysis by Streptococcus spp. On blood agar after 24 h of incubation at 37 ◦C. Figure S2 shows the appearance of Gram-negative rods under observation at the 100× lens of the microscope. At the same time, Figure S3 shows the appearance of Gram-positive cocci. Figure S4 shows the positive bile esculin test, which is for Enterococcus faecalis. Figure S5 shows the positive results for Escherichia coli using the API kit.

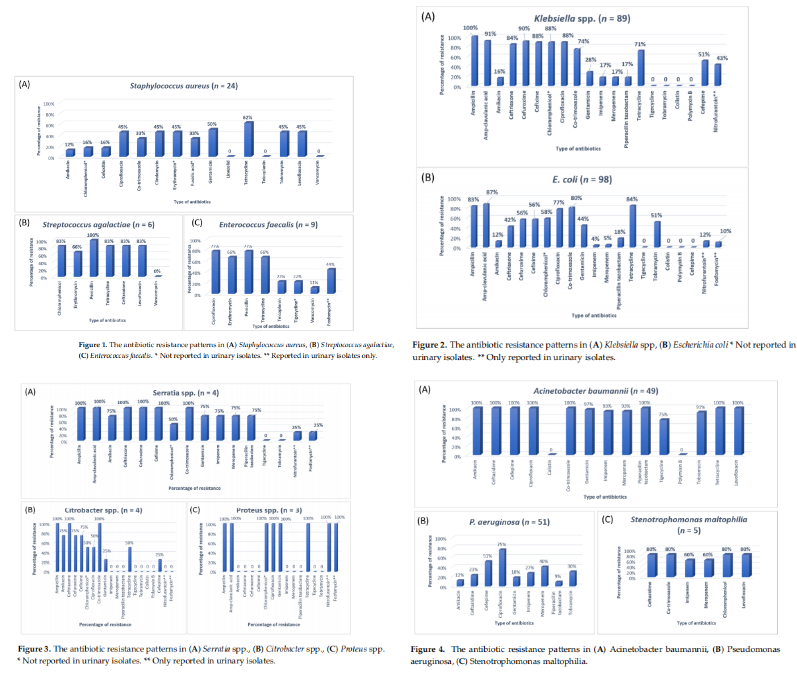
3. Antibiotic Susceptibility Patterns of Clinical Isolates
The antibiotic resistance patterns of individual bacterial isolates are shown in Figures 1–4.
In SICUs, different antibiotics were prescribed and given to patients with a different diagnosis, as shown in Table 3.

4. Distribution of COVID-19 Patients with Major Comorbidities
Apart from COVID-19, 36.09% (n = 309) of patients also suffered from bacterial pneumoniae, UTIs, meningoencephalitis and sepsis. The patients with pneumoniae were also diagnosed with aspiration, diabetes mellitus (DM), chronic obstructive pulmonary disease (COPD), ischemic heart disease (IHD) and hypertension (HTN). The patients with UTIs, meningoencephalitis and sepsis were also diagnosed with other comorbidities, which were the main reason for mortality, as shown in Table 4.

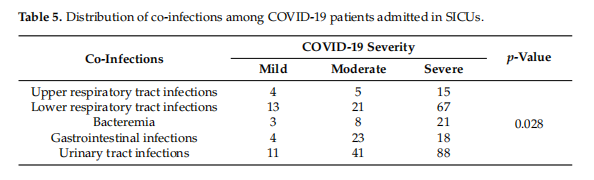
A significant association (p < 0.001) was found between the COVID-19 patients, comorbidities and the mortality rate. The correlation between co-infections and COVID-19 severity has been shown in Table 5.
5. Mortality and Recovery Rate of the Patient
The recovery and death rate among the COVID-19 patients was recorded during their stay in the hospital. From the total 856 COVID-19 patients, the recovery rate was 76.16% (n = 652). The death rate was 23.83% (n = 204). The most common death rate among COVID-19 patients with different comorbidities was the CLD cases, followed by pneumoniae, UTIs, sepsis, cellulitis, pancreatic cancer, ascites, CKD, pyelonephritis rigours, cystitis and hematuria.

Click here to get the Cistanche's benefits
Discussion
Antibiotics are the most commonly prescribed drugs among hospitalised patients, especially in SICUs. It is imperative to use the appropriate antibiotics in intensive care units with few prescriptions as an acceptable quality of care, infection control, cost reduction, and length of hospital stay [17–19]. Patients admitted in the SICU are critically ill requiring prescribed the medicine without waiting for the culture reports that give information about the antimicrobial resistance pattern of the suspected organism for a specific cause [20,21]. In terms of the culture reports, there is no possibility to wait for reports because these take a minimum of 48 h, and as a result antibiotic resistance occurs in patients admitted to the SICUs [22]. The current study was conducted among COVID-19 patients admitted in SICUs of tertiary care hospitals, requiring monitoring and special care to analyse the antibiotics utilisation pattern and determine the prevalence of AMR. Our analysis included each organism’s antibiotics sensitivity and resistance pattern isolated from patients admitted in SICUs.
A previous study on microbial infection and antibiotic resistance patterns in COVID-19 patients admitted in SICUs of tertiary care hospitals showed that Pseudomonas was the most common organism identified in the medical ICU, followed by Klebsiella pneumonia [23]. A study on the prevalence of microorganisms and bacterial resistance in the SICU of the Bangabandhu Sheikh Mujib Medical University of Bangladesh showed that the maximum identified organism was Acetobacter. (45.4%), of P. aeruginosa (32.2%), Proteus (11%), Klebsiella pneumoniae 10%, and E. coli (3%) were identified [24]. A study by Mehta et al., 2015 on ICU patients revealed that the Pseudomonas spp. (29.1%) was the most common organism, followed by Acinetobacter spp. (27.5%) [18]. Another previous analysis of AST and bacteriology profile on patients at tertiary care hospitals in Ahmadabad showed that Acinetobacter spp. [30.9%] was the most common organism, after coming Klebsiella spp. (29.8%) and P. aeruginosa (22.9%) [19]. However, in the current study, the most common isolated organism was E. coli (38%), followed by Klebsiella pneumoniae (24%), P. aeruginosa (14%). While Streptococcus agalactiae, Citrobacter freundii, Serratia liqeuficiens, and Stenotrophomonas maltophilla were 1.7%, 1.1%, 1.1% and 1.4%, respectively.

Cistanche extract and Cistanche powder
In Jordan, a study was conducted to see the AMR rates, the prevalence of bacterial infections, and misuse of antibiotics. They conducted a study to resolve the resistance rate of Gram-negative bacteria in patients admitted to the ICU of Prince Hasen hospital. It revealed that P. aeruginosa was the most resistant pathogen and broad-spectrum antibioticresistant bacteria [15]. A similar previous study on the uropathogens of ICU showed that E. coli was highly susceptible to imipenem, meropenem, and nitrofurantoin [25]. The second most common organism in the current study was Klebsiella pneumoniae, 100% resistant to ampicillin and 91% to Amp-clavulanic acid. For Klebsiella pneumoniae, the treatment option was amikacin and gentamicin. Work on the AMR pattern of the bacterial pathogen in the ICU of Fatimawati hospital showed high resistance to cephalexin (96.3%), cefotaxime (64.3%), and ceftriaxone (61.0%) shown by P. aeruginosa. The most effective antibiotic against P. aeruginosa was amikacin after imipenem (81.3%) and meropenem (75.2%). Klebsiella pneumoniae showed resistance to cephalexin, ceftriaxone, ceftazidime (86.6%) (75.9%) (73.4%) respectively [26]. In the current study, the most common organism, E. coli, showed ampicillin resistance (83%) and Amp-clavulanic acid (87%). In that case, the most effective antibiotic was imipenem and meropenem. Most E. coli showed high sensitivity to the imipenem and meropenem (96%). Imipenem and meropenem are usually used against both GPIs and GNIs. However, a study conducted in India showed that Klebsiella was more susceptible to imipenem and nitrofurantoin and showed more resistance to penicillin and gentamicin [18].
Most of the Gram-positive organisms in the current study were resistant to penicillin and tetracycline. For the treatment of the Gram-positive bacteria, linezolid, gentamicin, and vancomycin were used. Most Gram-negative organisms were resistant to two or more alarming antibiotics and shortly caused high mortality and morbidity. This will also affect the management of Gram-negative bacteria. Around 17 different types of antibiotics were used in the ICU. Carbapenem, fluoroquinolones, aminoglycoside, and quinolones were highly consumed among all the antibiotics. In a study in India, the high consumption of beta-lactam, carbapenem, metronidazole was observed. Another study in Nepal showed that ampicillin, metronidazole, and amoxicillin were the most common antibiotics [27]. Another research study showed that beta-lactam, nitroimidazoles, and fluoroquinolones were commonly used in an ICU [12]. The prevalence of MRSA in the current study was 16%, VRE was 11%, and CRE 17%, showing a high rate of AMR against most effective drugs. These high rates of AMR in SICU patients are an alarming situation for all of the clinicians and researchers and the time to take some actions to ensure the correct usage of antibiotics.

Herba Cistanche
Results of the current study showed that approximately 36% of COVID-19 patients had pneumonia followed by aspiration, diabetes mellitus (DM), chronic obstructive pulmonary disease (COPD), ischemic heart disease (IHD) and hypertension. UTIs, meningoencephalitis and sepsis followed by different comorbidities were also diagnosed. COVID-19 with bacterial pneumonia were the most important cause of distress and mortality. DM, HTN, CKD were significant comorbidities among ICU patients. Many patients had DM and HTN at the same time. Females experienced more complications and comorbidities than males. However, a previous systematic review and meta-analysis by Yang et al. (2020) reported that HTN and diabetes were the most common comorbidities, followed by cardiovascular diseases (CVD) and pneumonia. The pooled odds ratio of HTN, pneumonia and CVD in severe patients was lower than in non-severe patients [28]. We looked at the incidence of comorbidities in COVID-19 infection patients and discovered that underlying conditions, such as HTN, pneumonia, DM, CKD and COPD, might be a risk factor for high AMR rates among COVID-19 patients.
The current study was performed in a single hospital, and only those patients recruited for the current study were admitted in the SICUs. The sample size was smaller due to the single centre/unit study. Furthermore, multicentral studies with a larger sample size are recommended.

Cistanche's effects on kidney
Conclusions
The current study points to a significant prevalence of bacterial infections and high AMR rates in COVID-19 patients, especially with specific comorbidities and complications, making it challenging to identify the priority of treatment groups and improve the care of these infections. These high rates of AMR may lead to high mortality rates; that is why a specific antibacterial usage policy of institutions and SICUs are needed to control this problem. Periodic audits are required to monitor compliance with these guidelines. In a hospital setup, antibiotic sensitivity and AMR patterns must be considered to direct the infection control expert or clinician to begin empirical antibiotics in severe cases. An intensive and systematic effort is needed to quickly classify high-risk patients and minimise the irrelevant use of antibiotics, one of the primary reasons for increased AMR rates in developing countries.
References
17. Ahmed, N.; Ali, Z.; Riaz, M.; Zeshan, B.; Wattoo, J.I.; Aslam, M.N. Evaluation of Antibiotic Resistance and Virulence Genes among Clinical Isolates of Pseudomonas aeruginosa from Cancer Patients. Asian Pac. J. Cancer Prev. APJCP 2020, 21, 1333–1338.
18. Mehta, T.; Chauhan, B.; Rathod, S.; Pethani, J.; Shah, P.D. Bacterilogical profile and drug resistance pattern of isolates of the patients admitted in medical intensive care unit of a tertiary care hospital in Ahmedabad. Med. Sci. 2015, 4, 222–225.
19. Barai, L.; Fatema, K.; Haq, J.A.; Faruq, M.O.; Ahsan, A.A.; Morshed, M.A.H.G.; Hossain, M.B. Bacterial profile and their antimicrobial resistance pattern in an intensive care unit of a tertiary care hospital of Dhaka. Ibrahim Med. Coll. J. 2010, 4, 66–69.
20. Lockhart, S.R.; Abramson, M.A.; Beekmann, S.E.; Gallagher, G.; Riedel, S.; Diekema, D.J.; Quinn, J.P.; Doern, G.V. Antimicrobial resistance among Gram-negative bacilli causing infections in intensive care unit patients in the United States between 1993 and 2004. J. Clin. Microbiol. 2007, 45, 3352–3359.
21. Liang, S.Y.; Kumar, A. Empiric antimicrobial therapy in severe sepsis and septic shock: Optimising pathogen clearance. Curr. Infect. Dis. Rep. 2015, 17, 36.
22. Ahmed, N.; Zeshan, B.; Naveed, M.; Afzal, M.; Mohamed, M. Antibiotic resistance profile in relation to virulence genes fimH, hlyA and usp of uropathogenic E. coli isolates in Lahore, Pakistan. Trop. Biomed. 2019, 36, 559–568.
23. Sanjana, R.; Shah, R.; Chaudhary, N.; Singh, Y. Prevalence and antimicrobial susceptibility pattern of methicillin-resistant Staphylococcus aureus (MRSA) in CMS-teaching hospital: A preliminary report. J. Coll. Med. Sci.-Nepal 2010, 6, 1–6.
24. Radji, M.; Fauziah, S.; Aribinuko, N. Antibiotic sensitivity pattern of bacterial pathogens in the intensive care unit of Fatmawati Hospital, Indonesia. Asian Pac. J. Trop. Biomed. 2011, 1, 39–42.
25. Tuem, K.B.; Desta, R.; Bitew, H.; Ibrahim, S.; Hishe, H.Z. Antimicrobial resistance patterns of uropathogens isolated between 2012 and 2017 from a tertiary hospital in Northern Ethiopia. J. Glob. Antimicrob. Resist. 2019, 18, 109–114.
26. Mohamad, N.A.; Jusoh, N.A.; Htike, Z.Z.; Win, S.L. Bacteria identification from microscopic morphology: A survey. Int. J. Soft Comput. Artif. Intell. Appl. (IJSCAI) 2014, 3, 1–12.
27. Phillips-Jones, M.K.; Harding, S.E. Antimicrobial resistance (AMR) nanomachines—Mechanisms for fluoroquinolone and glycopeptide recognition, efflux and/or deactivation. Biophys. Rev. 2018, 10, 347–362.
28. Yang, J.; Zheng, Y.; Gou, X.; Pu, K.; Chen, Z.; Guo, Q.; Ji, R.; Wang, H.; Wang, Y.; Zhou, Y. Prevalence of comorbidities in the novel Wuhan coronavirus (COVID-19) infection: A systematic review and meta-analysis. Int. J. Infect. Dis. 2020, 10, 91–95.
Basit Zeshan 1 , Mohmed Isaqali Karobari 2,3,4, Nadia Afzal 5 , Amer Siddiq 6 , Sakeenabi Basha 7 , Syed Nahid Basheer 8 , Syed Wali Peeran 9 , Mohammed Mustafa 10 , Nur Hardy A. Daud 11 , Naveed Ahmed 1,12 , Chan Yean Yean 12 and Tahir Yusuf Noorani 2.
1. Department of Microbiology, Faculty of Life Sciences, University of Central Punjab, Lahore 540000, Pakistan; dr.basitzeshan@ucp.edu.pk (B.Z.); naveed.malik@student.usm.my (N.A.)
2. Conservative Dentistry Unit, School of Dental Sciences, Universiti Sains Malaysia, Health Campus, Kubang Kerian, Kota Bharu 16150, Kelantan, Malaysia; tahir@usm.my
3 Department of Conservative Dentistry & Endodontics, Saveetha Dental College & Hospitals, Saveetha Institute of Medical and Technical Sciences University, Chennai 600077, Tamil Nadu, India
4 Department of Restorative Dentistry & Endodontics, Faculty of Dentistry, University of Puthisastra, Phnom Penh 12211, Cambodia
5 Basic Health Unit Hospital (BHU) Mora, Tehsil and District Nankana Sahib, Nankana Sahib 39100, Pakistan; nadia.afzal511@gmail.com
6 Faculty of Medicine, Riphah International University, Islamabad 46000, Pakistan; 5400@students.riphah.edu.pk
7 Department of Community Dentistry, Faculty of Dentistry, Taif University, P.O. Box 11099, Taif 21944, Saudi Arabia; sakeena@tudent.edu.sa
8 Department of Restorative Dental Sciences, College of Dentistry, Jazan University, Jazan 45142, Saudi Arabia; syednahidbasheer@gmail.com
9 Department of Periodontics, Armed Forces Hospital Jizan, Jazan 82722, Saudi Arabia; doctorsyedwali@yahoo.in
10 Department of Conservative Dental Sciences, College of Dentistry, Prince Sattam Bin Abdulaziz University, P.O. Box 173, Al-Kharj 11942, Saudi Arabia; ma.mustafa@psau.edu.sa
11 Faculty of Sustainable Agriculture, Universiti Malaysia Sabah, Sandakan Campus, Locked Bag No.3, Sandakan 90509, Sabah, Malaysia; nur.hardy@ums.edu.my
12 Department of Medical Microbiology and Parasitology, School of Medical Sciences, Universiti Sains Malaysia, Kubang Kerian, Kota Bharu 16150, Kelantan, Malaysia; yychan@usm.my






