ACE1 Rs1799752 Polymorphism Is Not Associated With Long-COVID Symptomatology in Previously Hospitalized COVID-19 Survivors
Sep 18, 2023
Dear Editor
Viral mechanisms of infection of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the agent responsible for the coronavirus disease, 2019 (COVID-19), suggest the involvement of surface receptor for S1 of the angiotensin-converting enzyme 2 (ACE2) and transmembrane protease serine-2 (TMPRSS2) receptors.1 Single studies have found that single nucleotide polymorphisms (SNP), e.g., ACE2 rs2285666, ACE2 rs2074192, or TMPRSS2 rs12329760, seem to be potentially associated with the severity of COVID-19; however, current evidence is still not conclusive.2 Another potential SNP that could be implicated in various clinical diseases (e.g., renal damage, pneumonia, ischemic stroke) and immunological reactions (i.e., cytokine storm) induced by SARS-CoV-2 is ACE1 rs1799752. 3 In fact, evidence suggests that the D allele of this SNP is associated with higher severity of SARS-CoV-2.4
Cistanche can act as an anti-fatigue and stamina enhancer, and experimental studies have shown that the decoction of Cistanche tubulosa could effectively protect the liver hepatocytes and endothelial cells damaged in weight-bearing swimming mice, upregulate the expression of NOS3, and promote hepatic glycogen synthesis, thus exerting anti-fatigue efficacy. Phenylethanoid glycoside-rich Cistanche tubulosa extract could significantly reduce the serum creatine kinase, lactate dehydrogenase, and lactate levels, and increase the hemoglobin (HB) and glucose levels in ICR mice, and this could play an anti-fatigue role by decreasing the muscle damage and delaying the lactic acid enrichment for energy storage in mice. Compound Cistanche Tubulosa Tablets significantly prolonged the weight-bearing swimming time, increased the hepatic glycogen reserve, and decreased the serum urea level after exercise in mice, showing its anti-fatigue effect. The decoction of Cistanchis can improve endurance and accelerate the elimination of fatigue in exercising mice, and can also reduce the elevation of serum creatine kinase after load exercise and keep the ultrastructure of skeletal muscle of mice normal after exercise, which indicates that it has the effects of enhancing physical strength and anti-fatigue. Cistanchis also significantly prolonged the survival time of nitrite-poisoned mice and enhanced the tolerance against hypoxia and fatigue.

Click on chronic fatigue
【For more info:george.deng@wecistanche.com / WhatsApp:8613632399501】
Although ACE2, TMPRSS2, and ACE1 receptors are highly expressed in the lungs and respiratory tract, these receptors are also present in several other tissues explaining the heterogeneity clinical presentation of COVID-19. Accordingly, it could be proposed that these receptors are potentially also associated with the presence of symptoms after the acute phase of the infection, i.e., long-term. Mariani et al. have recently observed that post-COVID symptoms are present for up to one year in a heterogeneous population.5 Additionally, a recent meta-analysis reported that post-COVID fatigue can be present in up to 51% of COVID-19 survivors.6 Our group has recently identified that ACE2 rs2285666, ACE2 rs2074192, TMPRSS2 rs12329760 and TMPRSS2 rs2070788 polymorphisms do not predispose for long-COVID symptoms in previously hospitalized COVID-19 survivors.7
We present here a secondary analysis, the genetic part of THE LONG COVID EXPERIENCE STUDY,7 of the potential association between ACE1 rs1799752 polymorphism and the presence of long-COVID symptomatology.

Briefly, unstimulated whole saliva samples were collected from 288 COVID-19 survivors who were hospitalized at three urban hospitals in Madrid (Spain) during the first wave of the pandemic (March 2020). All participants provided written informed consent. The study was approved by the Institutional Ethics Committees of all involved institutions (HSO25112020; URJC0907202015920; HUFA20/126; HUIL/092-20). Genotyping collection and procedures have been previously published.7 Identification of each possible genotype of ACE1 rs1799752 polymorphism was conducted by using specific fluorescent dyes. The ACE1 rs1799752 is an insertion/deletion of an Alu repeat sequence that is interrogated using a pair of assays. The possible variants of the ACE1 rs1799752 (the mutant Alu insertion allele -I allele- and the wild type of deletion allele -D allele-) SNP lead to the following genotypes (D/D, D/I, I/I) derived from the sequence:
CCCATTTCTCTAGACCTGCTGCCT [-/ALU] ATACAGTCACTTTTATGTGGTTTC

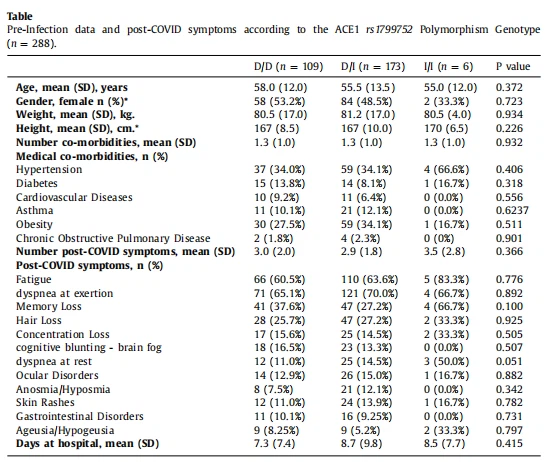
Demographic, medical comorbidities, and hospitalization data were collected from hospital medical records. Post-COVID symptomatology was collected during a face-to-face appointment conducted by experienced healthcare professionals at 17.8 (SD 5.2) months after hospitalization.7 Dyspnea at exertion (n = 196, 68%) and fatigue (n = 181, 63%) were the most prevalent post-COVID symptoms in the total sample. Other post-COVID symptoms included pain (n = 117, 40.6%), memory loss (n = 92, 32%), or hair loss (n = 77, 26.7%). The genotype distribution (D/D genotype n = 109, 37%; D/I genotype n = 173, 60%; I/I genotype n = 6, 3%) deviated from that expected based on the Hardy-Weinberg equilibrium, a result also extensively found in published COVID-19 studies. Overall, no significant differences in long-COVID symptoms were observed depending on the genotypes of ACE1 rs1799752 SNP (Table). No sex differences in the genotype distribution (P = 0.723) were either found.
The results of this secondary analysis reveal that ACE1 rs1799752 polymorphism does not predispose to developing long-term symptoms when evaluated in previously hospitalized COVID-19 survivors. Current results would agree with our previous report on ACE2 and TMPRSS2 polymorphisms.7 The lack of association between these SNPs and long-COVID could be explained by the significant gene variability existing (i.e., ethnic differences) across populations.8 Further, the lack of relationship among SNPs associated with COVID-19 severity and the presence of long-COVID reinforce the idea that the severity of the disease is not associated with the development of post-COVID symptoms since the presence of these symptoms is similar between hospitalized and nonhospitalized patients.9 Similarly, the D allele of the ACE1 rs1799752 polymorphism has been associated with a higher risk of hospitalization in patients with co-morbidities at the acute phase of the infection since these individuals are at a higher risk for developing acute respiratory distress syndrome (ARDS).10 In our study, no differences in previous medical co-morbidities depending on the ACE1 rs1799752 genotype were observed.
Some limitations of the current data should be recognized. First, only previously hospitalized COVID-19 survivors were included, therefore, the role of these SNPs in non-hospitalized patients is yet to be investigated. Second, it could be possible that larger samples could identify genotype differences, accordingly, our data should be considered exploratory. Population-based cohort studies and a whole genome SNP analysis might help validate current results and identify other genes potentially related to long-term symptoms.
Role of the funding source
The project was supported by a grant from Novo Nordisk Foundation NNF21OC0067235 (Denmark) and by a grant associated to the Fondo Europeo De Desarrollo Regional - Recursos REACT-UE del Programa Operativo de Madrid 2014–2020, en la línea de actuación de proyectos de I+D+i en materia de respuesta a COVID 19 (LONGCOVID-EXP-CM). Both sponsors had no role in the design, collection, management, analysis, or interpretation of the data, draft, review, or approval of the manuscript or its content. The authors were responsible for the decision to submit the manuscript for publication, and the sponsor did not participate in this decision.
Declaration of interests
No conflict of interest is declared by any of the authors
Acknowledgments
The Center for Neuroplasticity and Pain (CNAP) is supported by the Danish National Research Foundation (DNRF121) and Norvo Nordic Foundation (NNF21OC0067235). We also thank the Genomics Unit, Madrid Science Park Foundation, Spain for its valuable support.

References
1. Singh HO, Choudhari R, Nema V, Khan AA. ACE2 and TMPRSS2 polymorphisms in various diseases with special reference to its impact on COVID-19 disease. Microb Pathog 2021;150:104621.
2. Gupta K, Kaur G, Pathak T, Banerjee I. Systematic review and meta-analysis of human genetic variants contributing to COVID-19 susceptibility and severity. Gene 2022;844:146790.
3. Adli A, Rahimi M, Khodaie R, Hashemzaei N, Hosseini SM. Role of genetic variants and host polymorphisms on COVID-19: from viral entrance mechanisms to immunological reactions. J Med Virol 2022;94:1846–65.
4. Aziz MA, Islam MS. Association of ACE1 I/D rs1799752 and ACE2 rs2285666 polymorphisms with the infection and severity of COVID-19: a meta-analysis. Mol Genet Genomic Med 2022;10:e2063.
5. Mariani C, Borgonovo F, Capetti AF, Oreni L, Cossu MV, Pellicciotta M, Armiento L, Bocchio S, Dedivitiis G, Lupo A, Galli M, Rizzardini G. Persistence of Long-COVID symptoms in a heterogenous prospective cohort. J Infect 2022;84:722–46.
6. Ji G, Chen C, Zhou M, Wen W, Wang C, Tang J, Cheng Y, Wu Q, Zhang X, Wang M, Feng Z. Post-COVID-19 fatigue among COVID-19 in patients discharged from hospital: a meta-analysis. J Infect 2022;84:722–46.
7. Fernández-de-las-Peñas C, Arendt-Nielsen L, Díaz-Gil G, Gómez-Esquer F, Gil-Crujera A, Gómez-Sánchez SM, Ambite-Quesada S, Palomar-Gallego MA, Pellicer-Valero OJ, Giordano R. Genetic association between ACE2 (rs2285666 and rs2074192) and TMPRSS2 (rs12329760 and rs2070788) polymorphisms with post-COVID symptoms in previously hospitalized COVID-19 survivors. Genes 2022;13:1935.
8. Smatti MK, Al-Sarraj YA, Albagha O, Yassine HM. Host genetic variants potentially associated with SARS-CoV-2: a multi-population analysis. Front Genet 2020;11:578523.
9. Fernández-de-las-Peñas C, Rodríguez-Jiménez J, Cancela-Cilleruelo I, Guerrero-Peral A, Martín-Guerrero JD, García-Azorín D, Cornejo-Mazzuchelli A, Hernández-Barrera V, Pellicer-Valero OJ. Post-COVID-19 symptoms 2 years after SARS-CoV-2 infection among hospitalized vs. non-hospitalized patients. JAMA Netw Open 2022;5:e2242106.
10. Sabater Molina M, Nicolás Rocamora E, Bendicho AI, Vázquez EG, Zorio E, Rodriguez FD, Gil Ortuño C, Rodríguez AI, Sánchez-López AJ, Jara Rubio R, Moreno– Docón A, Marcos PJ, García Pavía P, Villa RB, Gimeno Blanes JR. Polymorphisms in ACE, ACE2, AGTR1 genes and severity of COVID-19 disease. PLoS One 2022;17:e0263140.
【For more info:george.deng@wecistanche.com / WhatsApp:8613632399501】






