Cloning, Functional Identification And Expression Analysis Of Chalcone Synthase From Cistanche Tubulosa
Mar 22, 2024
Abstract: Objective
To study the function of chalcone synthase from Cistanche tubulosa and the expression pattern of CtCHS gene in white, red and purple flowers.
Methods The CtCHS gene was cloned by RT-PCR and bioinformatics analysis was carried out. The pET-28a-CtCHS plasmid was transferred into the Escherichia coli to induce protein expression by IPTG. The CtCHS recombinant protein was incubated with the substrates p-coumaroyl CoA and malonyl CoA, and the products were detected by HPLC and MS. The pCAMBIA1300-35S-GFP-CtCHS plasmid was transferred into the protoplast of Arabidopsis thaliana to observe the subcellular localization of the CtCHS protein. The expression pattern of CtCHS gene in the flowers of C. tubulosa with three colors was detected by qRT-PCR.
Results A CtCHS gene was cloned. The length of the open reading frame (ORF) was 1173 bp, encoding 390 amino acids, and the molecular weight of the protein was 42 8000.

ORGANIC CISTANCHE EXTRACT WITH 30% ECHINACOSIDE AND 12% ACTEOSIDE
CtCHS contained the catalytic residues Cys164, His303, Asn336, which were conserved in chalcone synthase. The CtCHS protein was expressed in E. coli and purified to obtain the recombinant protein. CtCHS protein could catalyze p-coumaroyl CoA and malonyl CoA to produce naringenin chalcone. CtCHS protein was mainly localized in the cytoplasm. CtCHS gene was highly expressed in purple and red flowers, and the relative expression levels were 8.44 and 3.21 times higher than that in w hite flowers, respectively.
Conclusion A CtCHS gene was cloned from the flower of C. tubulosa and protein was expressed. CtCHS protein has chalcone synthase function, and the expression of CtCHS gene is higher in darker flowers, indicating that CtCHS gene may regulate the flower color of C. tubulosa, which lays a foundation for further research on the regulation mechanism of the flower color of C. tubulosa.
Key words: Cistanche tubulosa (Schenk) Wight; chalcone synthase; enzyme activity analysis; subcellular localization; expression analysis
Cistanche Tubulosa (SCHENK) WIGHT is the genus Pyreal plant, which is naturally distributed in southern Xinjiang [1] in the Hetian area of Xinjiang [2]. The pipe flowers are high in medicinal value. Its meat stems are one of the sources of the 2020 version of "Chinese Pharmacopoeia". [3] Research on the pipe flowers cordonus meat stems has increased, but there are relatively few biological research on its flowers. The flowering period of the pipe flowers is from April to May [4], mainly There are three colors of white, red, and purple. The pipe flowers are rich in color, but their flower color regulation mechanism has not been reported. Classes are related to the type, content, content, and distribution of petal tissue structure, pigment, and regulation of environmental factors and genetic factors [5]. The pigment in plant flowers mainly includes three categories: flavonoids, carotene, and beetine. Color, the most widely Great color range [7]. The types and content of flavonoids affect the color of many plants, such as the three varieties of Paeonia Lactiflora Pall.
It appears as white, and the main color of the color contains the color of the red and beautiful competence makes the flowers present as purple -red [8]. Floral color is an important feature of many ornamental plants, and it is also the key factor that attracts the pink insects [9]. Studies have shown that it is passed down The highest frequency of insect visits is the deep purple cistanche plant [10]. For biological research on the synthetic genetic research of cistanoids in the tube flower cistanosaurus flowers, it is conducive to clarifying the tube flower cordon coloring control mechanism, providing theoretical basis for improving the color of genetic engineering technology, promoting its richer colors, increasing viewing, expanding applications scope.
The biosynthetic pathway of flavonoids begins with the phenylpropanoid pathway. Phenylalanine is first converted into cinnamic acid under the action of phenylalanine ammonialyase (PAL), and cinnamic acid is 4-hydroxylated by cinnamic acid. The enzyme (cinnamic acid-4-hydroxylase, C4H) and 4-coumarate coenzyme A ligase (4-coumarate coenzyme A ligase, 4CL) catalyze the reaction to generate 4-coumaroyl-CoA [11]. One molecule of 4-coumaroyl-CoA and three molecules of malonyl-CoA are catalyzed by chalcone synthase (CHS) to generate naringenin chalcone, which has flavonoid compounds .The basic skeleton of chalcone isomerase, flavanone-3-hydroxylase, dihydroflavonol 4-reductase, etc. generates a variety of flavonoids such as isoflavones, flavonols, anthocyanins, etc. [12]. As a key enzyme in the flavonoid biosynthetic pathway, CHS regulates the content of flavonoids in plants and also plays an important role in regulating plant flower color [13]. For example, post-transcriptional gene silencing of Gentiana scabra Bunge CHS can inhibit the biosynthesis of anthocyanins and cause specific whitening of corolla lobes [14]. CHS gene silencing of Actinidia eriantha Benth. CHS reduces the anthocyanin content in petals and causes The color of the red petals becomes lighter [15]. Therefore, in this study, a CtCHS gene was cloned from Cistanche deserticola flowers, and bioinformatics analysis, prokaryotic expression and purification, enzyme activity analysis, subcellular localization analysis and expression analysis in three colors of flowers were performed on it to explore The function of CtCHS protein and the expression pattern of CtCHS gene were used to preliminarily explore the relationship between CtCHS and Cistanche deserticola flower color, which laid the foundation for the study of the regulatory mechanism of Cistanche deserticola flower color.
1 Materials and instruments
1.1 Materials
Cistanche tuberosa flowers were collected from Yutian County, Hotan Prefecture, Xinjiang in May 2018. The random principle was followed during sampling, and Cistanche tuberosa flowers with good physiological status and consistent plant height were selected. Three strains of Cistanche deserticola with three flower colors of white, red, and purple were taken and identified as Cistanche tubulosa (Schenk) Wight by Professor Tu Pengfei of Peking University School of Pharmacy. They were quickly frozen in liquid nitrogen, stored on dry ice, and transported to Beijing. E. coli DH5α chemically competent cells were purchased from Nanjing Novezan Biotechnology Co., Ltd., and E. coli BL21 (DE3) chemically competent cells were purchased from Beijing Quanshijin Biotechnology Co., Ltd.; pET-28a, pCAMBIA1300-35S-GFP The vector was purchased from Wuhan Transduction Biology Laboratory Co., Ltd. Malonyl-CoA was purchased from Sigma Company, 4-coumaroyl-CoA, naringenin-chalcone were purchased from Shanghai Yuanye Biotechnology Co., Ltd., and naringenin was purchased from Shanghai McLean Biochemical Technology Co., Ltd.
1.2 Instruments
Nanodrop 2000C spectrophotometer (Thermo Fisher Company, USA); gradient PCR instrument (Eppendorf Company, Germany); CFX 96 fluorescence quantitative PCR instrument, UVP gel imager, gel electrophoresis instrument, protein electrophoresis instrument (BioRad Company, USA) ;EnSpire
Microplate reader (PerkinElmer Company, USA); constant temperature incubator (Shanghai Jinghong Experimental Equipment Co., Ltd.); DHZ-D large-capacity full-temperature oscillator (Taicang Experimental Equipment Factory); THZ-82A water bath constant temperature oscillator ( Jiangsu Kexi Instrument Co., Ltd.); AIRTECH ultra-clean workbench (Suzhou Antai Air Technology Co., Ltd.); high-pressure steam sterilizer (TOMY Company, Japan); Milli Q pure water instrument (Millipore, United States); LC-20A high-efficiency liquid Phase chromatograph (Shimadzu Company, Japan); UPLC-Q-Exactive-Orbitrap high-resolution mass spectrometry (Thermo Fisher Company, USA); FV1200 laser confocal microscope (Olympus Company, Japan).

2 Experimental methods
2.1 RNA extraction and cDNA synthesis. Cistanche deserticola flowers were ground in liquid nitrogen and then subjected to RNA extraction assay.
Total RNA was extracted using a kit (Norgen, Cat 17200), the RNA concentration and quality were detected by NanoDrop 2000C, and RNA samples with an A260/A280 ratio between 1.8 and 2.1 were selected for reverse transcription synthesis of cDNA. M-MLV reverse transcriptase (Promega, M170B) was used to reverse-transcribe the qualified total RNA into cDNA. The experimental operations were performed according to the instructions.
2.2 Cloning of CtCHS gene
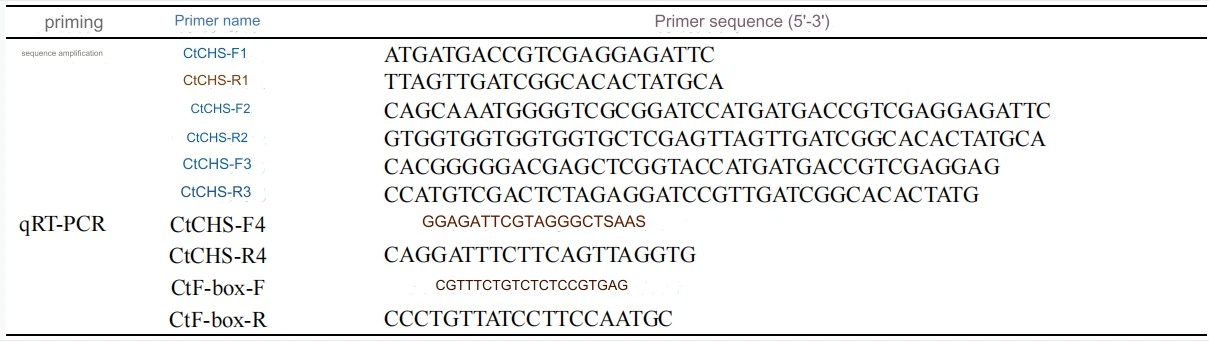
Based on the Cistanche deserticola flower transcriptome data of our research group [16], a CtCHS with a complete open reading frame was screened out, and SnapGene software was used to design specific primers based on the gene sequence (Table 1). PCR amplification was performed using Cistanche deserticola flower cDNA as a template. The amplification system was 2×Phanta Max Buffer 25 μL, dNTP Mix (10 mmol/L) 1 μL, upstream and downstream primers (10 μmol/L) 2 μL each, cDNA 2 μL, Phanta Max Super- Fidelity
DNA Polymerase 1 μL, ddH2O 17 μL. The reaction conditions were: pre-denaturation at 95°C for 3 min; 25 cycles of denaturation at 95°C for 15 s, annealing at 55°C for 15 s, and extension at 72°C for 1 min; and reaction extension at 72°C for 5 min. The PCR product was detected by 1% agarose gel electrophoresis, and the DNA purification kit (Novizan, Cat DC301-01) was used for gel recovery, and then the recovered DNA was purified by a rapid cloning kit (Novizan, Cat C601-01). The DNA fragment was ligated with the pCE2 TA/Blunt-Zero vector, transformed into Escherichia coli DH5α competent cells, and after culturing overnight at 37°C, single clone strains were selected and sent to the company for sequencing.
Table 1 Primer sequences
2.3 Bioinformatics analysis
Use ProtParam online software to analyze protein physical and chemical properties; use Prot Scale online tool to analyze protein hydrophilicity/hydrophobicity; use online tool SOPMA to predict protein secondary structure; use online analysis software SWISS-MODEL to predict protein tertiary structure; use online software TMHMM Predict the transmembrane structure of the protein; use DNAMAN software to compare the homology of CtCHS with the CHS amino acid sequences of other species; use MEGA-X software to construct a phylogenetic tree, using neighbor-joining (NJ) and Poisson correction The evolutionary distance is calculated using the Bootstrap method, and the number of Bootstrap repetitions is 1000 times.

UPLC-Q-Exactive-Orbitrap MS was used for further detection. The liquid phase conditions were Waters Acquity UPLC BEH Sheild RP18 column (100 mm × 3.0 mm, 1.7 μm), column temperature 40 ℃, naringenin, chalcone and naringenin. The loading volume of the control substance is 4 μL, and the loading volume of the enzyme reaction product is 7 μL. Water (A)-acetonitrile (B) is used as the mobile phase. The elution gradient is: 0 to 5 min, 30% B; 5 to 15 min. , 30%~60% B; 15~20 min, 60%~100% B; 20~25 min, 100% B; 25~31 min, 100%~30% B. The volumetric flow rate is 0.3 mL/min. The mass spectrometry conditions are electrospray ion source, nitrogen as carrier gas, sheath gas pressure 3.5 MPa, auxiliary gas pressure 1.0 MPa, spray pressure 3500 V, capillary temperature 350°C, auxiliary gas heating temperature 200°C, positive and negative ion modes, scanning ion range m/z is 100~1500, Full MS resolution is 35 000, dd-MS2 resolution is 17 500, (N)
CE/stepped nce set to 35, 60.
2.6 Subcellular localization of CtCHS protein
Based on the CtCHS sequence and the principle of seamless cloning, primers with enzyme cutting sites Kpn Ⅰ and BamH Ⅰ were designed, and the corresponding target fragments were amplified by PCR. The pCAMBIA1300-35S-GFP plasmid was digested with restriction endonucleases Kpn Ⅰ and BamH Ⅰ, and the target fragment and vector were connected through the seamless cloning kit (Novizan, Cat C115-01), and then transferred into E. coli DH5α competent cells. , select single clone strains for sequencing, and extract plasmids from the correct strains. The pCAMBIA1300-35S-GFP-CtCHS and pCAMBIA 1300-35S-GFP plasmids were transformed into Arabidopsis thaliana protoplasts using PEG4000, and cultured under low light for 8 to 10 h. The subcellular localization of CtCHS protein was observed under a laser confocal microscope.

2.7 CtCHS expression pattern analysis
Based on the CtCHS sequence, DNAMAN was used to design real-time fluorescence quantitative primers, with F-box as the internal reference gene. The primer sequences are shown in Table 1. The relative expression of CtCHS in Cistanche deserticola flowers of different colors was detected by real-time fluorescence quantitative PCR. Use TransStart Green qPCR SuperMix (full gold, AQ101-02) to measure with SYBR Green fluorescent dye method, reaction system: 2×TransStart Green qPCR SuperMix 5 μL, upstream and downstream primers (10
μmol/L) 0.2 μL each, cDNA template 0.5 μL, nucleotidase-free water 4.1 μL, a two-step method was used for the reaction, and the reaction procedure was based on the method of Dong Xianjuan et al. [18]. Each sample contained 3 biological replicates, and the experiment was repeated three times. Experimental data were analyzed using Excel to 2
The relative expression of CtCHS was calculated using the −ΔΔCt method, and GraphPad Prism 8 was used to perform significance analysis and draw histograms.