Innate And Adaptive Immune Abnormalities Underlying Autoimmune Diseases: The Genetic Connections(1)
Dec 28, 2023
Except for an extremely small number of cases caused by single gene mutations, most autoimmune diseases result from the complex interplay between environmental and genetic factors. In a nutshell, the etiology of the common autoimmune disorders is unknown despite progress elucidating certain effector cells and molecules responsible for pathologies associated with inflammatory and tissue damage. In recent years, population genetics approaches have greatly enriched our knowledge regarding genetic susceptibility of autoimmunity, providing us with a window of opportunities to comprehensively re-examine autoimmunity-associated genes and possible pathways. In this review, we aim to discuss the etiology and pathogenesis of common autoimmune disorders from the perspective of human genetics. An overview of the genetic basis of autoimmunity is followed by 3 chapters detailing susceptibility genes involved in innate immunity, adaptive immunity, and inflammatory cell death processes respectively. With such attempts, we hope to expand the scope of thinking and bring attention to lesser-appreciated molecules and pathways as important contributors of autoimmunity beyond the 'usual suspects' of a limited subset of validated therapeutic targets.

cistanche plant-increasing immune system
autoimmune diseases, etiology, pathogenesis, innate immunity, adaptive immunity
Introduction
Autoimmunity refers to a wide array of collective conditions in which broken self-tolerance leads to manifestations of pathological alterations and clinical symptoms resulting from immune responses against self-targets. One of the major challenges of treating autoimmune conditions is the almost impossible mission of reversal of break of tolerance. When patients walk into rheumatology clinics, they often complain about swollen joints or morning stiffness whereas few would state the emergence of anti-nuclear antibodies or high levels of rheumatoid factors as the main reason for their doctor visits. Consequently, rheumatologists face the challenges of treating illnesses whose etiology stems from uncharacterized events that may be traced back for an extended period, sometimes decades ago. The current therapeutic strategies for autoimmune diseases are analogous to the solutions practiced by firefighters, i.e., putting forward the best attempts to extinguish the fire regardless of the cause that originally ignited the flammables.
Information that may shed light on the etiology of autoimmunity is in part provided by the fact that certain individuals are more susceptible to given diseases than the rest of the general population. Analyses of genetic predispositions using population genetics approaches such as genomewide association studies (GWAS) have yielded rich information regarding the etiology and pathogenesis of autoimmunity. Albeit many susceptibility loci are located in the non-coding segments of the genome with unclear functional implications, a substantial proportion of the described loci fall into protein-coding regions that encode products involved in a wide variety of biological processes. While a subset of autoimmune susceptibility genes presents themselves with clear ties to immune phenotypes given the well-established roles in innate and/or adaptive immune responses, connections of other genes such as those whose functions are predominantly implicated in developmental processes and metabolic activities to autoimmunity are more cryptic. In this extensive review, we discuss what we have learned from the recent studies aiming to elucidate the underlying genetic components of autoimmunity and how such knowledge may educate us toward a better understanding of autoimmunity-associated immune abnormalities. In combination with progressively deepened knowledge of contributing environmental factors, the ultimate goal is to provide insights into the development of novel therapeutic approaches against autoimmunity and to alleviate symptoms or even achieve disease-free status for autoimmune patients.

cistanche tubulosa-improve immune system
The genetic basis of autoimmune diseases
Genetic epidemiology of autoimmune disease
Autoimmune diseases are one of the most common diseases worldwide and have a significant public impact because of their high morbidity and mortality (Rioux and Abbas, 2005). The general prevalence of autoimmune diseases ranged from less than 5 per 100,000 (e.g., uveitis (Miserocchi et al., 2013), Wegener granulomatosis (Cotch et al., 1996)) to more than 500 per 100,000, such as rheumatoid arthritis (RA) (Almutairi et al., 2021) and ankylosing spondylitis (AS) (Dean et al., 2014). Although most autoimmune diseases can occur at any age, the peaks of onset differ by illness (Amador-Patarroyo et al., 2012). For instance, type 1 diabetes (T1D) (Maahs et al., 2010) primarily occurs in childhood and adolescence, but multiple sclerosis (MS) (Schwehr et al., 2019) and systemic lupus erythematosus (SLE) (Mina and Brunner, 2013) mostly appear during the mid-adult years, and RA (Symmons, 2002) mainly among older people. In addition, autoimmune diseases present gender disparities with a greater prevalence amongst women in a 2:1 ratio (Angum et al., 2020). Furthermore, the genetic epidemiology of autoimmune diseases becomes more complicated when variations in ethnicity, geographical regions, and susceptibility genes are considered (Wang et al., 2015). Coeliac disease is a typical example, which is less prevalent in Asia. This may be due to the genetic factor that carriers of the HLA-DQ2 antigens linked to celiac disease occur in 5%– 10% of Chinese and sub-Saharan Africans when compared to 5%–20% in Western Europe. In contrast, HLA-DQ8 occurs in 5%–10% of English, Tunisians, and Iranians, but less than 5% in Eastern Europeans, Americans, and Asians (Kang et al., 2013). Collectively, autoimmune diseases are common diseases with genetic heritability and exhibit gender and age disparities with ethnic and geographic differences (Cooper and Stroehla, 2003). The genetic heritability of autoimmune disease varies greatly (Ramos et al., 2015), for instance, from very high in AS (>90%) (Brown et al., 2016) to relatively low in inflammatory bowel disease (IBD, 12%) and MS (15%) (Kuusisto et al., 2008), whereas RA and SLE have a mid average genetic heritability of roughly 60% (Guerra et al., 2012). The explanation for these differences is mainly due to the heritability and the interaction result of epigenetic factors and environmental factors (Baranzini and Oksenberg, 2017). Furthermore, the hereditary bias between familial cases and the general population for prevalent complicated diseases should be taken into account (Momozawa et al., 2018). Mounting evidence indicates a tendency toward familial aggregation of autoimmune disease (Cárdenas-Roldán et al., 2013). Many family studies have indicated that first-degree relatives (FDRs) of individuals with diagnosed autoimmune disease (e.g., RA, MS, AS) had increased familial risk of acquiring certain other autoimmune disease versus control probands (Cooper et al., 2009), which is even higher in monozygotic twins (Bogdanos et al., 2012). Furthermore, researchers also observed that not only FDRs but also spouses of individuals with autoimmune diseases are at increased risk (Emilsson et al., 2015).
Genetic factors of autoimmune diseases
The GWAS revolution has accelerated the identification of autoimmune disease-associated variants
Detecting the exact disease-causal variants and susceptibility loci may help us to better understand the mapping between genotype and phenotype of complex traits in disease mechanisms (Hirschhorn et al., 2002). However, the traditional linkage analysis methods are not enough to map genomic loci accurately due to the great variability and extensive linkage disequilibrium (LD) of the majority of autoimmune diseases (Fernando et al., 2008). The GWAS revolution in the early 2000s, which is a powerful tool for unbiasedly identifying regions of the genome related to human variation and disease, opened up new avenues for global research into the inheritance patterns of autoimmune disease (Visscher et al., 2012). The joint Genome-wide association study coordinated by the Wellcome Trust Case Control Consortium (WTCCC) in 2007 was the first real advance step forward in discovering new genetic correlations of autoimmune disease susceptibility through GWAS (Wellcome Trust Case Control Consortium, 2007). Notably, this WTC study revealed several novel genes that are highly linked to RA, T1D, and celiac disease (a type of IBD). For the first time, researchers have uncovered a gene called PTPN2 that links these three autoimmune diseases. In the same year, WTC published another large-scale genetic study in AS and MS, reporting two new AS loci: ARTS1 and IL23R, and highlighting that IL23R may as a shared susceptibility factor for the major 'seronegative' diseases like AS and Crohn's disease (CD) and psoriasis (Wellcome Trust Case Control Consortium and The Australo-Anglo-American Spondylitis Consortium (TASC), 2007).
Growing international collaboration across different ethnic cohorts expands the sample size of GWAS studies, yielding more compelling autoimmune disease findings. As an example, the International Genetics of Ankylosing Spondylitis Consortium (IGAS) consortium conducted a dense SNP genotyping study in 10,619 cases and 15,145 controls of European, East Asian, and Latin American ancestry in 2013, which increased the number of AS-associated loci to 31 (including 13 new loci) and 12 additional AS-associated haplotypes at 11 loci, revealed the critical role of aberrant peptide processing before major histocompatibility complex (MHC) class I presentation and alterations of the IL-23 proinflammatory cytokine pathway in the pathogenesis of AS (International Genetics of Ankylosing Spondylitis Consortium et al., 2013). In addition, a meta-analysis of GWAS is becoming increasingly prominent, which can improve the ability to identify association signals by combining samples from multiple cohorts to detect more variations and cover more genomic areas than a single dataset (Zeggini and Ioannidis, 2009). In 2014, A three-stage trans-ethnic meta-analysis (Okada et al., 2014) conducted for 100,000 subjects of European and Asian ancestries (29,880 RA cases and 73,758 controls) discovered 42 novel RA risk loci and identified 98 biological candidate genes in a total of 101 risk loci. Particularly, they devised an in silico pipeline developed by well-established bioinformatics methodologies and based on functional annotation to identify 98 biological candidate genes at these 101 risk loci, and first conducted functional annotation of RA risk SNPs which aim to provide empirical evidence for drug discovery. Furthermore, employing the cross-disease meta GWAS methodology can improve the power to identify common susceptibility loci across several autoimmune diseases, even if the association signals differ between diseases. As early as 2011, Zhernakova et al. (Zhernakova et al., 2011) discovered fourteen non-HLA common loci associated with the mechanism of antigen presentation and T-cell activation in a meta-analysis of two published GWAS on celiac disease and RA of European ancestry cohorts. Similarly, cross-disease meta GWAS analysis for five published data of psoriasis and Crohn's disease performed by Ellinghaus et al. (Ellinghaus et al., 2012) identified 20 shared disease association loci and tested cross-disease associations in additional cohorts in 2012. Then in early 2019, the first cross-disease genome-wide meta-analysis in systemic seropositive rheumatic illnesses (including systemic sclerosis, SLE, RA, and idiopathic inflammatory myopathies) (Acosta-Herrera et al., 2019) was published, which revealing five new shared genome-wide significant independent loci from a cohort of 11,678 patients and 19,704 non-affected controls from European ancestry groups.

cistanche benefits-strengthen immune system
Post-GWAS era: the functional genomics of autoimmune disease
However, while GWAS has successfully led to the discovery of thousands of loci that have been statistically related to disease and trait risk, several challenges and limits have emerged. Including biological relevance to disease and clinical utility for prognosis or treatment lag far behind, could not explain the vast majority of genetic heredity for diseases, as well as the research constraint at the cell level (Visscher et al., 2017). Naturally, the post-GWAS era has arrived, which is primarily arguing for causation and risk gene identification (Pierce et al., 2020), as well as encouraging the transition from association to function (Gallagher and Chen-Plotkin, 2018).
During this period, numerous remarkable studies with newly-rising so-called post-GWAS methods were published. One of the most effective techniques is "fine mapping", for identifying trait-relevant genetic elements in a genomic locus that has already been restricted, which has proven useful in translating GWAS findings into possible therapies (Schaid et al., 2018). A new method known as CC-GWAS (case-case genome-wide association study) (Peyrot and Price, 2021) has recently gained traction. Peyrot W.J. et al. use summary statistics from the respective case-control GWAS to test for variations in allele frequency between cases of two disorders, which transcends conventional approaches that require individual-level information. They have effectively identified loci with varying allele frequencies among patients of eight psychiatric diseases via CC-GWAS and validated the CCGWAS method using three publicly available autoimmune disease GWAS datasets, which include CD, ulcerative colitis (UC), and RA. Demonstrate the ability to use this strategy to improve clinical diagnoses and treatment of other autoimmune diseases. Furthermore, population cohort studies also play a critical role in proving causality and promoting medication development (Wijmenga and Zhernakova, 2018). Recently, a large-scale meta-analysis across East Asian and European populations on RA via various post-GWAS approaches performed by Eunji Ha et al. (Ha et al., 2021) integration of accumulated knowledge of RA variants with emerging high throughput omics data which led to the identification of 11 new RA susceptibility loci. So far, enormous success of wide-scale genetic studies in identifying genetic variants of autoimmune disease. The question of how to make effective use of these data remains a challenge.
The shared genetic mechanisms between autoimmune diseases
The previously epidemiological studies have demonstrated that human autoimmune diseases are complex disorders that result from the interaction between genic susceptibility and environmental factors (Wang et al., 2015). Even though autoimmune diseases are heterogeneous conditions in clinical and therapeutic features with the involvement of multiple organ systems (Ramos et al., 2015), it is the consensus that autoimmune diseases share complicated and similar genetic backgrounds. Meanwhile, genetic research also supports the existence of distinct pathogenesis pathways for various autoimmune diseases (Richard-Miceli and Criswell, 2012). A decade ago, researchers already found almost half of the 107 immune disease-risk SNPs across seven immune-mediated inflammatory and autoimmune diseases (including CD, MS, psoriasis, RA, SLE, and T1D) are shared, which as is the case with alleles in the major histocompatibility locus (Cotsapas et al., 2011).
Lately, Caliskan M. et al. (Caliskan et al., 2021) developed a catalog of 85 fine mapping studies on autoimmune GWAS locus by combining text mining with a systematic review. They compiled 230 GWAS loci which consist of 455 combinations of locus-by-disease association signals with 15 autoimmune diseases. To refine the genes shared by the main autoimmune diseases in this study, we chose five major autoimmune disorders (CD, RA, T1D, IBD, MS) to display the overlapping genes within these diseases with the information of disease association loci (Figure 1A). The 74 GWAS loci spanning more than two primary autoimmune diseases from this catalog were then visualized using a chord diagram based on the causative gene confidence scores of these GWAS loci and their associated autoimmune disorders (Figure 1B). Notably, we can see that IL2 and TAGAP are both found shared in four autoimmune diseases (IL2 in CD, IBD, RA, and T1D, and TAGAP in IBD, MS, RA, and T1D, respectively), which is consistent with earlier mice studies and clinical experiment results (Chen et al., 2020; Clough et al., 2020; Pérol et al., 2016). The insight gained from these analyses underlined the crucial function of regulatory T cells in immunological homeostasis and will contribute to the development of regulatory T-cell treatments for these autoimmune diseases.

cistanche benefits for men-strengthen immune system
Click here to view Cistanche Enhance Immunity products
【Ask for more】 Email:cindy.xue@wecistanche.com / Whats App: 0086 18599088692 / Wechat: 18599088692
Many common variations have been discovered in autoimmune diseases and other human disorders, so-called "pleiotropy" (Inshaw et al., 2018). It is widely established that immune system dysfunction is linked to an increased risk of Parkinson's disease (PD) (Tan et al., 2020). For example, in a comprehensive epidemiological study of 310,000 people in Sweden with 33 different autoimmune diseases, the prevalence of increased risk of Parkinson's disease was as high as 33% (Li et al., 2012). In contrast, findings from a large GWAS involving 47,580 instances of RA and 482,703 cases of PD suggest that Rheumatoid arthritis lowers the risk of Parkinson's disease (Li et al., 2021a). The association between autoimmune diseases and Parkinson's disease appears to be inconclusive, which could be related to sample size constraints. Nevertheless, the study of pleiotropy between autoimmune diseases and other comorbidities could aid in the discovery of novel loci not previously associated with the disease. Witoelar et al. reported that 17 novel loci were identified as the overlap between PD and autoimmune diseases including the 4 known PD loci (GAK, HLA-DRB5, LRRK2, and MAPT) which presented in RA, UC, and CD, and highlighted the involvement of Human leukocyte antigen (HLA) (Witoelar et al., 2017).
Gene-environment interaction effect on autoimmune disease
It is consensus that environmental factors and ascertainment bias may play a role in disease risk in addition to genetic factors. With the development industrial civilization advances and science and technology advances, including new industries, new chemicals, and pharmaceuticals. The prevalence of autoimmune diseases caused by environmental factors has increased. Improving the understanding of "environmental triggers" to autoimmune diseases may help people avoid dangers and determine treatment alternatives (Gioia et al., 2020; Vojdani, 2014). In recent years, an increasing number of studies have demonstrated that smoking (Ishikawa and Terao, 2020), red meat (Pattison et al., 2004), and the high-sodium diet (Salgado et al., 2015) have negative consequences on disease development, whereas a vegetarian diet (Kjeldsen-Kragh et al., 1991), polyunsaturated fatty acids (Fetterman Jr. and Zdanowicz, 2009), vitamin D (Jeffery et al., 2016), and probiotics (Bungau et al., 2021) contribute to improved health assessment. Consequently, dietary patterns and supplements were encouraged as future adjuvant therapy in the treatment of autoimmune disease, such as the Mediterranean Diet (MD), which consists primarily of vegetables, fruit, fish, olive oil, and dairy products (Pocovi-Gerardino et al., 2021).

Figure 1 Overlapping genes (GWAS loci) association between main autoimmune diseases. A, The UpSet graphic generated by UpSetR (Conway et al., 2017) shows the number of GWAS loci that overlap for each of the five common autoimmune diseases (CD, Crohn's disease; RA, rheumatoid arthritis; T1D, and type 1 diabetes; IBD, inflammatory bowel disease; MS, multiple sclerosis), which is based on the comprehensive catalog of autoimmune disease GWAS fine-mapping research. B, The chord diagram created with the R package "circle" (Gu et al., 2014), presents the relationship between 74 GWAS loci (left) that overlap in more than two major autoimmune diseases and the diseases with which they are associated (right).
Modulation of gut microbiota-derived metabolites is one of the most important indirect mechanisms of how dietary habits and nutrition influence disease progression (Han et al., 2021). Altered microbiota composition has been linked to reduced intestinal barrier function and mucosal immune system dysregulation (Khan and Wang, 2019), one of the well-known hypotheses is "the gut-joint axis" (Zaiss et al., 2021). However, it is uncertain if gut dysbiosis is a cause or an effect of autoimmune disease. A result from the general population cohort of all babies in the Southeast Sweden project indicated that genetic risk for T1D autoimmunity is related to unique alterations in the gut microbiota (Russell et al., 2019). Evidence from animal and human studies indicated that HLA alleles influence the process of the gut microbiota interacting with host immunity (Xu and Yin, 2019). Besides, an emerging study using shotgun metagenomics on AS patients indicates an enrichment of potentially cross-reactive bacterial epitopes, and the TNFi therapy has an effect on microbiome composition (Yin et al., 2020).
Advanced genetic study and perspective of autoimmune disease
The GWAS successfully applied to the identification of a large number of genetic variants (mainly SNPs) of the disease and the association with many complex traits. However, it had limited predictive ability in diseases. The polygenic risk score (PRS) profiling method, which can aggregate the effects of variants across the genome, is able to be used in the estimation of an individual's genetic liability to a trait or disease by calculating based on disease genotype profile and relevant GWAS data. Li Z. et al. (Li et al., 2021b) highlighted the significant diagnostic capacity of PRS in AS patients compared to traditional diagnostic test methods including C-reactive protein (CRP), HLA-B27, and sacroiliac MRI. For reality clinical application, more study of PRS applied in autoimmune diseases within specific ethnic groups is required. In 2020, Choi et al. (Choi et al., 2020) published a tutorial on performing polygenic risk score analyses in nature protocols calculated according to their genotype profile and relevant GWAS data, which may help in the interpretation of PRS-trait associations.
Another challenge of GWAS is that it is difficult to confirm causal genes or disease distal regulatory areas that relate to cell type-specific behaviors. Single-cell RNA sequencing (scRNA-Seq) is a powerful method for collecting gene expression in individual cells from living tissues using high throughput sequencing analysis across the entire transcriptome. It has already shown promising results in MS, AS, and RA. For example, recent research by Simone D. et al. (Simone et al., 2021) applied single-cell transcriptome analysis by using peripheral blood and synovial fluid samples from patients with AS and psoriatic arthritis (PsA), with results showing detailed characterization of Tregs cells and demonstrated LAG-3 directly inhibits IL-12/23 and TNF secretion by patient-derived monocytes, which could be a potential mechanism for SpA. The majority of current autoimmune disease treatments rely on systemic immunosuppression, which makes patients prone to infections. Precision medicine is regarded as the cornerstone of future cancer therapies (Shin et al., 2017), involving the development of novel diagnostics and customized drugs to a patient's individual needs based on genetic, biomarker, phenotypic, or psychological characteristics. In mid-2016, Ellebrecht C. T. et al. (Ellebrecht et al., 2016) showed that chimeric antigen receptor T cells (CAR-T cells) can be modified to seek out and kill self-reactive B cells, which may provide specific targeting of autoreactive B cells in antibody-mediated autoimmune disease and eventually help identify potential therapies.
Nowadays, the intersection of precision medicine and artificial intelligence (especially machine learning and deep learning algorithms) is a popular area in autoimmune disease medical research. The goal is to better capture individual variation in genes, function, and environment in order to build and optimize diagnostic, therapeutic, and prognostic pathways (Jameson and Longo, 2015). This could lead to a new chance to personalize therapies specifically for autoimmune disease patients, particularly those with rare autoimmune diseases, in order to achieve effective treatment (Subramanian et al., 2020). Meanwhile, the ethical and legal issues surrounding artificial intelligence-driven healthcare are gaining traction in society and provoke discussion (Amann et al., 2020).
Genetic components underlying innate immune abnormalities in autoimmune diseases
Autoreactive immune responses are the major pathogenic driving force in autoimmune diseases. Although different autoimmune diseases manifest themselves with drastically distinct symptoms, certain common genetic risk factors broadly underlie autoimmunity-associated immune abnormalities (Cho and Feldman, 2015). In the following three chapters, based on the knowledge obtained from human genetics studies such as GWAS, we discuss the plausible factors and pathways that might be involved in aberrant activation of the immune system. The first part of this chapter briefly discusses risk genes with apparent connections to innate immune functions while the next two chapters focus on T cell-related immune abnormalities and pathogenic factors in TNF signaling. In addition, in the second and third parts of this chapter, we expand the discussion of the risk genes whose connections to autoimmunity may not appear as obvious at first glance, exemplified by those genes and pathways best known for their roles in developmental and metabolic processes.
Autoimmune disease susceptibility genes with well-characterized functions in innate immunity Risk genes in the antigen presentation process
In homeostasis, DCs are the major inducer of peripheral tolerance for humoral immunity, in which tolerogenic DCs induce depletion or energy in auto-reactive T cells and also polarize the T cells into regulatory T cells (Iberg et al., 2017). While, in autoimmunity activation, bypassed peripheral tolerance could result from the genetic risk variants in antigen presentation pathways and/or hyper-activation of T cells (Theofilopoulos et al., 2017). With extensive investigations, HLA risk variants in autoimmune diseases are related to the abnormal antigen presented by DCs, and non-HLA risk factors could also be involved. The autoimmune disease-related polymorphisms in ERAP1 and ERAP2 loci have been linked to impropriate antigen presentation, resulting from the disturbed antigen peptide trimming by ERAP1/2 encoded enzymes for MHC-I presentation. Additionally, the T cells hyper-proliferation and inflammatory polarization could be genetically influenced by autoimmune disease-causative risk alleles, such as PTPN22 for TCR signaling, IL12A and STAT4 for Th1 polarization by IL-12, and IL23R for IL-23 mediated Th17 polarization.
Pathogenic interaction between plasmacytoid DCs (pDCs) and neutrophils
One characteristic of a subset of autoimmune diseases represented by SLE is the increase of pDCs in circulation and in disease-affected tissues, such as the kidney (Coutant and Miossec, 2016). pDCs are the professional type I interferon (IFN) producing cells, which play an essential role in antiviral innate immune response. In SLE-related conditions, accumulated pDCs in circulation and local lesion tissues promote inflammation and autoantibody production, largely dependent on type I IFNs (Soni and Reizis, 2019). In SLE pathogenesis, a devastating forward loop is mediated by neutrophils, pDCs, and B cells. In inflammatory environments, such as SLE patients' renal tissues, neutrophils are recruited and activated by inflammatory cytokines, such as IL-8 and IL-17 (Fresneda Alarcon et al., 2021). Upon activation, neutrophils could undergo suicidal cell death, NETosis, which releases DNA contents from the neutrophil extracellular trap (NET). The NET structure contains nuclear DNA and oxidated mitochondrial DNA, both of which are potent TLR9 agonists and auto-antigens for type I IFN production in pDCs and autoantibody production in B cells, respectively (Soni and Reizis, 2019). In such pDC-mediated interferonopathy, multiple signaling components are genetically pre-deposited to disease-prone conditions (Mohan and Putterman, 2015). Firstly, the polymorphism in genes related to TLR signaling pathway components has been identified in autoimmune diseases, such as IRAK1 and IRF5. For predisposed TLR signaling in SLE pathogenesis, resulting in type I IFN production in pDC has been considered as a major player, and crosstalk between TLR7/9 signaling and B cell activation has also been implied in the promotion of autoantibody-producing plasma cells (Suthers and Sarantopoulos, 2017). Some gene loci for signaling regulators are also identified as risk alleles, such as TNFAIP3 and TNIP3 for NF-κB signaling that contribute to the inflammatory phenotypes in myeloid cells. Secondly, the abnormal type I IFN signaling could also be genetically imprinted in autoimmune disease patients, where the genes encoding type I IFN receptor and downstream signaling cascade kinase, IFNAR1 and TYK2, have been identified with autoimmune disease-related polymorphism. In contrast to SLE, pDC function in other autoimmune diseases is less characterized, and the less commonly shared pDC-mediated pathogenesis has been implied among different diseases. Similarly to SLE, in mouse models for type I diabetes, pDC has been shown to promote disease progression by producing type I IFN (Reizis, 2019). While, tolerogenic pDC phenotypes in synovium and periphery of RA patients have been described (Cooles et al., 2018; Kavousanaki et al., 2010; Takakubo et al., 2008), and such protective role is supported by the worsened disease conditions with pDC depletion in an arthritis mouse model (Jongbloed et al., 2009).
Monocyte and macrophage-mediated inflammation
The central role of macrophages and monocytes in autoimmune disease-related tissue inflammation is underpinned by the success of therapeutic interventions targeting inflammatory cytokines produced by monocytes and macrophages (Conigliaro et al., 2019). In general, inflammatory macrophages and monocytes are considered effector cells for disease-related tissue inflammation (Navegantes et al., 2017). With functional dissection for commonly presented risk alleles across different autoimmune diseases, the genetically pre-deposited pathogenic pathways in macrophages and monocytes have been unraveled. In such inflammatory responses, myeloid cells, specifically the monocyte-derived macrophages, are the central node to sense the pro-inflammatory environments resulting in inflammatory mediator production and further bridge the local adaptive immune activation by favoring the differentiation of pathogenic CD4+ T cells and autoantibody-secreting plasma cells (Tsokos, 2020; Weyand and Goronzy, 2021).
Immune complex-induced inflammatory responses in inflamed tissues are universally manifested by Fc receptor signaling activation and complement pathway activation in myeloid cells, and the anti-complement therapy and blockade of Fc receptor exhibit promising clinical efficacy in RA and SLE patients (Galindo-Izquierdo and Pablos Alvarez, 2021; Zuercher et al., 2019). Genetic variants in both complement and Fc receptor pathways are also associated with autoimmune diseases (Theofilopoulos et al., 2017). In particular, polymorphisms in ITGAM and FCGR2B loci have been correlated with impaired negative regulation of immune complex-induced inflammation. ITGAM encodes CD11b, also known as complement receptor CR3, to mediate complement-dependent phagocytosis for immune complex and apoptotic cells, and C3b activates CR3 to induce the production of anti-inflammatory cytokines in macrophages. For polymorphism in the FCGR2B locus, the impaired function of FcγRIIB-mediated repression on Fc receptor signaling could result in excessive inflammatory activation of myeloid cells by the immune complex. Thus, the accumulation of immune complex and cell debris and downstream hyper-inflammatory activation of macrophages in local lesion tissues are genetically pre-deposited in autoimmune diseases to promote pathogenesis. Given the genericity of immune complex accumulation presented in autoimmune disease settings, such abnormally activated Fc receptor signaling, and complement pathways are considered to be therapeutically targeted with broad effects in different diseases.
Although the shared genetic risk factors and inflammatory myeloid cells could be observed under various autoimmune diseases, the responsiveness of different autoimmune disease patients to myeloid-targeting therapy could be totally different. For anti-GM-CSF therapy, RA patients receive prominent disease control from therapy, but disease conditions were worsened by anti-GM-CSF intervention in SLE patients (Lotfi et al., 2019). The mechanism underlying such difference is still unclear, but it suggests that myeloid cells receiving GM-CSF signaling in SLE patients could be protective. This practical issue suggests that acknowledging the heterogeneity of inflammatory conditions helps the development of anti-inflammatory therapy by targeting the critical inflammatory mediators in specific disease settings. Macrophages and monocytes could be over-activated by a plethora of inflammatory stimuli in autoimmune diseases, which further results in systemic inflammation termed macrophage activation syndrome (Crayne et al., 2019). Such systemic inflammation mediated by macrophages and monocytes could develop global symptoms and even be life-threatening. Macrophage activation syndrome could be observed in systemic juvenile idiopathic arthritis (sJIA), where the pro-inflammatory phenotypes are presented in the periphery, as blood monocytes of patients produce an array of inflammatory cytokines, such as TNF, IL-6, IL-1β. In such hyper-inflammatory macrophage activation, the dysregulated inflammatory pathways in innate immunity could result from genetic variants, such as IRF5 for type I IFN signaling, NLRC4 for IL-1β-producing inflammasome pathway, and TNFAIP3 for NF-κB signaling (Schulert and Cron, 2020).
Notch signaling is genetically associated with autoimmune diseases
As previously mentioned, genetic risk factors associated with self-tolerance and inflammatory pathways contribute to autoimmune diseases. Meanwhile, genetic variants that could not be directly aligned to abnormalities of immune responses are being functionally annotated with accumulating knowledge (Figure 2), within which we will discuss the pathological processes participated by Notch signaling and mitochondria-centric metabolism, two pathways with genetic relevance to autoimmune diseases. Notch signaling is one of the essential pathways for embryonic tissue and organ development, after which Notch signaling also maintains local homeostasis in various tissues. In mammals, four Notch signaling receptors (Notch1–4) and five Notch ligands have been identified. Upon activation by Notch ligands, the receptor proteolytically releases the Notch intracellular domain (NICD) and regulates gene expression via interaction with RBPJ which is the central nuclear transcription regulator in the canonical Notch signaling pathway. Notch signaling is broadly related to the immune system, where Notch signaling participates in the development of immune cells in both lymphoid and myeloid lineages and also regulates the function of terminally differentiated immune cells to fine-tune the immune responses in homeostasis and disease conditions (Vanderbeck and Maillard, 2021). In recent studies, accumulating evidence on Notch signaling-involved pathogenesis in multiple autoimmune diseases suggests targeting Notch-related pathways would be a promising therapeutic intervention. Therefore, a summary of current knowledge on the regulation of pathogenesis, especially the inflammatory responses, in autoimmune diseases by the Notch pathway is informative and insightful for the design of therapy.
Three out of four Notch receptor gene loci have been identified as risk alleles in autoimmune diseases (Table 1). In addition, the gene locus of RBPJ has been identified as an RA risk allele, and loci of DLL1 and DLL4 have been identified as risk alleles in SLE, multiple sclerosis, and type I diabetes (Table 1). Thus, the dysregulated Notch signaling is genetically implied in autoimmune diseases. Herein, we will discuss the Notch-related regulation during the pathogenesis of RA and SLE, two autoimmune diseases associated with risk variants in Notch receptors, ligands, and RBPJ loci with high prevalence and broad attention.

Figure 2 Functional category of genetic components underlying autoimmune diseases
Upregulated Notch signaling contributes to RA pathogenesis
In RA patients, Notch receptor activation has been identified in both immune and non-immune cells. In lymphoid lineage, activation of Notch1 has been identified in synovial T cells (Yabe et al., 2005), and the peripheral T cells of active RA patients show upregulated Notch2, 3, 4 expression and Notch signaling activation (Jiao et al., 2010). In collagen II-immunized mice, the Notch signal is activated in the synovium, and RA-like manifestations are alleviated upon Notch signaling inhibition by γ-secretase inhibitors (Choi et al., 2018; Jiao et al., 2014; Jiao et al., 2011; Park et al., 2015). In most of these studies, the altered Th1/Th17 to Treg ratio by the Notch signaling pathway was suggested for progressive disease outcomes in mouse models (Choi et al., 2018; Jiao et al., 2014; Jiao et al., 2011), where Notch3 and DLL1 promote both Th1 and Th17 expansion (Jiao et al., 2011), DLL3 promotes Th17 expansion (Jiao et al., 2014), and Notch1 suppresses Treg population (Choi et al., 2018). However, the context-dependent Notch signaling function presented by different studies could be the result of variations among different experiment systems. Although the promotion of Th17 expansion by DLL1 was shown by Jiao et al. via DLL1 treatment of in vitro splenic mononuclear cells (Jiao et al., 2011), in another study, DLL1 failed to promote Th17 expansion (Jiao et al., 2014). Therefore, whereas the beneficial effects of blocking Notch signaling were consistent in RA disease mouse models, the detailed mechanism, especially the specific contributions from each Notch receptor and ligand remain to be clarified. Lineage-specific knockout mice for Notch receptors and ligands should be useful to specifically evaluate Notch signaling function in T cell-mediated RA pathogenesis.
Notch signaling activation is also observed in myeloid cells in both RA patients and RA disease mouse models (Sekine et al., 2012; Sun et al., 2017). As mentioned in RA pathogenesis, myeloid cells, specifically monocytes, and monocyte-derived cells, could function either as macrophages or osteoclasts to promote inflammation and bone erosion. While, Notch signaling, indeed, participates the both parts, modulating the polarization of macrophages and differentiation of osteoclasts. Notch signaling promotes inflammatory polarization of macrophages under various conditions (Shang et al., 2016). However, the proinflammatory function of Notch signaling in macrophages is much less characterized in RA-related inflammation. Indeed, global inhibition of notch signaling results in the reversed hyper-inflammatory phenotype in macrophages in the RA disease mouse model (Sun et al., 2017). As discussed in Notch function in RA T cells, it is also important to investigate the specific contribution of Notch receptor-ligand pairs in macrophages to the inflammatory response in RA pathogenesis. For osteoclastogenesis, Notch signaling has been shown to both positively and negatively regulate osteoclast differentiation (Shang et al., 2016), which could be the result of either different receptor-ligand Notch signaling activation or the variations in different experimental systems. Actually, the regulation by different Notch receptor-ligand pairs in RA-related osteoclastogenesis could be hypothesized, as a previous study showing that Notch2/DLL1 promote, but Notch1/Jagged1 suppress osteoclast development under a unified experimental system (Sekine et al., 2012). Notch signaling also plays an important role in synovial fibroblast-mediated RA pathogenesis. The pathogenesis contributed by Notch-mediated pathogenic fibroblasts is relatively well documented, as most findings are derived from patient samples, and were solidified by a well-controlled in vitro system. Firstly, the Notch activation in synovial fibroblast was identified in RA patients' synovium (Ando et al., 2003; Ishii et al., 2001; Nakazawa et al., 2001a; Nakazawa et al., 2001b; Wei et al., 2020; Yabe et al., 2005). In addition, activated Notch signaling is induced by inflammatory environments, such as excessive TNF and hypoxia (Ando et al., 2003; Gao et al., 2015; Gao et al., 2012; Jiao et al., 2012; Nakazawa et al., 2001a; Nakazawa et al., 2001b). Inhibition of Notch activation by γ-secretase inhibitor not only alleviated inflammation-induced fibroblast proliferation but also decreased inflammatory cytokine production, such as IL-6 (Jiao et al., 2012; Nakazawa et al., 2001a). For specific mechanisms, Notch1 signaling has been functionally implicated in inflammatory RA fibroblasts (Gao et al., 2015; Nakazawa et al., 2001a; Nakazawa et al., 2001b), and Notch3 signaling has been recently identified to mediate the pathogenic expansion of THY1-expressing RA fibroblasts (Wei et al., 2020). Given the successful attempts in experimental animal models, the Notch signaling pathway may represent a promising therapeutic target of RA. While the detailed mechanism, especially the Notch signaling regulated Th1/Th17 response in RA pathogenesis, still needs to be further investigated. Moreover, for a better therapy design, the evaluation of RA pathogenesis that is mediated by different Notch receptors or ligands should be performed comprehensively, as evidenced by only moderate disease alleviation by blocking Notch1 compared to Notch3 blockade in K/BxN mice serum transferring RA mouse model (Wei et al., 2020).
Table 1 Summary of Notch receptors, ligands and RBPJ-related risk alleles identified by GWAS studies in autoimmune diseases
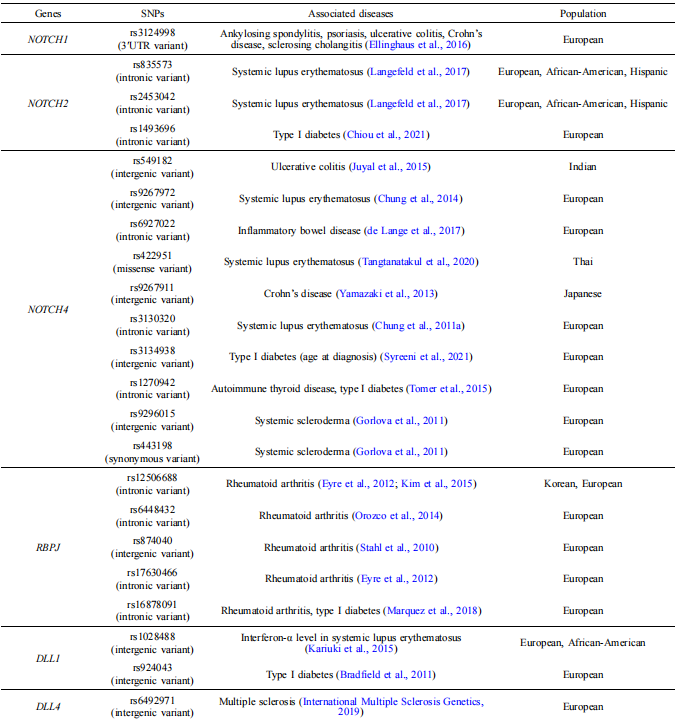
Dysregulated Notch signaling is associated with SLE pathologies
In SLE patients, upregulation of Notch signaling has been observed in locally damaged renal tissues. Specifically, Notch3 expression is upregulated in kidney tissues of patients with lupus nephritis compared to healthy individuals (Breitkopf et al., 2020). Moreover, the activation of Notch signaling in renal tissues with lupus nephritis is evidenced by the cleavage of Notch1 and Notch2 and the nuclear localization of Notch1 and Notch3 in podocytes (Lasagni et al., 2010; Murea et al., 2010). In addition to patients, the experimental mouse model showing lupus-like syndromes also exhibits altered Notch signaling in renal tissues (Breitkopf et al., 2020; Lemos et al., 2019; Zhang et al., 2010). Although the global Notch signaling inhibition alleviates the damage of renal tissues and autoantibody production in a lupus mouse model (Teachey et al., 2008; Zhang et al., 2010), the contributions from Notch signaling in specific cell types to the SLE pathogenesis are still under evaluation. On one hand, Notch signaling is required for mounting adequate Th1 response during T-cell activation, and Th17 differentiation has also been substantially proven to be dependent on Notch signaling (Tindemans et al., 2017). On the other hand, Treg cells could be negatively regulated by Notch signaling in mouse models for autoimmune diseases (Tindemans et al., 2017). As mentioned in the RA disease setting, the pro-inflammatory role of Notch signaling in T cell response could be extrapolated from the current knowledge that elevation of Notch signaling in RA T cells accounts for the enhanced Th1 and Th17 but downregulated Treg response. In SLE patients, Notch signaling in T cells is dysregulated in the opposite directions between local damaged tissues and peripheral blood. In contrast to the upregulation of Notch signaling in damaged renal tissues (Breitkopf et al., 2020), the evidence showing decreased Notch signaling in peripheral T cells was collected as, first, the deceased Notch1 expression in SLE patients' T cells is mediated by cAMP-responsive element modulator α (CREMα) associated suppressive epigenetic regulation (Rauen et al., 2012); second, the Notch signaling activation could also be downregulated in SLE patients' T cells via soluble CD46 interfered interaction between Jagged1 and Notch receptor (Ellinghaus et al., 2017). Although inhibition of Notch signaling with γ-secretase inhibitor in lupus-prone mice significantly alleviates lupus-related autoantibody production, nephritis, and local inflammation, the Notch signaling mediated T cell response seems to play a protective role in SLE disease progression. In SLE patients and lpr mice, disease-associated guanidinylated YB-1 functions as a ligand for Notch3 to potently mediate Notch activation in kidney tissues and promote IL-10 production in T cells via Notch3 (Breitkopf et al., 2020). Similarly, in SLE peripheral blood, interfered Notch activation by soluble CD46 impedes the switching of IFN-γ+ Th1 cells to IL-10 producing IFN-γ+ Th1 cells (Ellinghaus et al., 2017), which have been named as Tr1 cells and play the role in peripheral immune tolerance (Pot et al., 2011). Meanwhile, the effector T cells in SLE patients are resistant to the suppressive regulation by Treg (Vargas-Rojas et al., 2008; Venigalla et al., 2008). Mechanistically, Notch activation in effector T cells potentiates receiving TGF-β signaling from Treg to mediate immune tolerance (Grazioli et al., 2017). Thus, bypass of suppression by Treg in SLE T cells is highly possible to be attributed to decreased Notchcoopted TGF-β signaling. Given the crucial role of Treg in immune tolerance, the quantitative and qualitative defects in Treg have also been identified in SLE patients (Valencia et al., 2007; Vargas Rojas et al., 2008). Then, could decreased Notch activity in T cells contribute to dysfunctional Treg response in SLE patients as well? Currently, the studies by specific manipulation of Notch signaling in mouse Treg cells suggest the suppressive function of Notch signaling in Treg maintenance or function in autoimmune disease-related settings (Charbonnier et al., 2015; Rong et al., 2016). However, the Notch-related regulation in Treg development and function is rather complex, exemplified by the inconsistent conclusions on Notch-mediated FOXP3 expression drawn by studies with various experimental systems, and the outcome of Notch activation during Treg development and maintenance depends on the contexts, the different combinations of Notch receptor-ligand pairs, different types of Treg cells, and specific tissue environments, which has been comprehensively reviewed by Paola G. et al. (Grazioli et al., 2017). For Treg in SLE patients, low CD25 expression is another disease-related Treg defect that could be related to Notch signaling (Horwitz, 2010). Although the IL2RA (CD25) gene locus has been identified as one of the risk alleles for SLE, such genetic mutation could barely explain the low CD25 expression in resting SLE T cells (Costa et al., 2017). Interestingly, Notch signaling could maintain CD25 expression in T cells (Adler et al., 2003), making it possible that downregulation of Notch signaling might lead to the defect of CD25 expression in SLE patients' T cells. Therefore, the decreased Notch activity in SLE patients' T cells could be hypothesized as one of the reasons for defects in Treg, and the further dissection of Notch signaling in SLE Treg cells would be informative to elaborate the notch-regulated Treg response during SLE pathogenesis.
According to the direct characterization of Notch signaling in kidney tissues of SLE patients, Notch signaling activation has been evidenced by increased cleavage of Notch1 and Notch2 and increased expression of Jagged1 in glomerulus (Murea et al., 2010), and increased nuclear Notch1 and Notch3 in PDX+ podocytes and increased nuclear Notch3 in CD24+ renal podocyte progenitor cells in renal tissues of SLE patients (Lasagni et al., 2010). Moreover, the severity of glomerulosclerosis is closely correlated to the increase of cleaved Notch1 in podocytes (Murea et al., 2010). Podocytes are the major structural component of the glomerulus, and lupus nephritis manifests the loss of podocytes. Pathologically, Notch signaling contributes to the loss of podocytes during pathogenic conversion in renal tissues by regulating podocyte regeneration. For detailed mechanisms, human podocyte progenitor cells were isolated and cultured to differentiate into podocytes in vitro, and the downregulation of Notch signaling is related to G2/M cell cycle arrest during differentiation (Lasagni et al., 2010). Although enforcing the Notch signaling activation by overexpressing Notch3-NICD in podocyte progenitor cells promotes the expression of podocyte marker genes, the progenitor cells were pushed through the cell cycle checkpoint by activated Notch signaling, which further resulted in cell death of podocytes by defected mitosis (Lasagni et al., 2010). In addition to lupus nephritis, other nephropathic models in mice also show that Notch signaling activation, specifically Notch1, and Notch3, in podocytes results in loss of podocytes, renal tissue damage, and impaired kidney function (Asanuma et al., 2017). However, Notch2 activation in podocytes seems to function as the protective feedback loop to constrain cell death (Asanuma et al., 2017). Given the increased Notch2 activation in podocytes of SLE patients (Murea et al., 2010), the global inhibition of Notch signaling by γ-secretase inhibitor might undesirably lead to the dysfunction of such intrinsic protective pathway. Unlike the overall pathogenic effect of upregulated Notch signaling in RA-related disease conditions, the disturbances of Notch signaling in SLE patients mediate different regulations to disease progression in context-dependent manners. Therefore, targeting Notch signaling for therapeutic purposes with global inhibition for activation is undesirable (Grosveld, 2009), and intervention with specificity for Notch signaling in SLE therapy is anticipated. Of note, DLL4 expression in DCs is upregulated under inflammatory conditions in both humans and mice, and DLL4 in DCs could activate Notch signaling in T cells to induce Th1 and Th17 responses, which suggests the potential therapeutic strategy by targeting DLL4 to reverse the Th1/Th17 dominant inflammatory response (Meng et al., 2016). Moreover, the expansion of thymic DCs by DLL4 blockade promotes Treg differentiation (Billiard et al., 2012), and inactivation of DLL4-mediated Notch signaling in autoimmune disease mouse models results in disease remission and less inflammatory T cell response (Billiard et al., 2012; Reynolds et al., 2011). As above, not only the genetic association, but the dysregulation of Notch signaling in autoimmune diseases leads to the disease progression in autoimmune diseases. Given receptors, ligands and even critical enzymes in Notch signaling are promising druggable targets, further translational investigations that target the context-dependent regulation by Notch, especially for T cells in autoimmune diseases, require a detailed evaluation for the specific Notch receptor-ligand pairs that contribute to the imbalanced Th1/Th17 autoimmune response. Last but not least, the actual genetic effects on Notch signaling are still largely unknown in autoimmune diseases. The majority of identified genetic variants aligned to Notch signaling are distributed in non-coding regions (Table 1), which suggests that these genetic variants mediate transcriptional regulation via DNA regulatory elements, but do not interfere with the translation. Therefore, the actual positive or negative regulation of the Notch signaling should be further evaluated with advanced methodologies, such as CRISPR-Cas system-implemented genome editing, and the cell-type specific regulation should also be considered during the evaluation (Stewart et al., 2020).
Mitochondria-centric metabolism-associated genes in autoimmunity
Among hundreds of risk loci identified in autoimmune diseases, there are genes related to mitochondrial function and cellular metabolism. For example, C4orf52 in RA and CMC1 in AS encode the components of the mitochondrial translation regulation assembly intermediate of cytochrome c oxidase (MITRAC) (Ellinghaus et al., 2016; Okada et al., 2014; Timón-Gómez et al., 2018). Such SNPs suggest the dysregulated electron transport in the inner membrane of mitochondria, and subsequent mitochondrial hyperpolarization and impaired ATP synthesis could lead to ROS production, metabolic program switching, and vulnerability to inflammation-induced cell death (McGarry et al., 2018). To discuss the pathogenic role of mitochondria malfunction in autoimmune diseases, we will focus on mitochondria-centric ROS production and metabolic disturbance during disease progression and metabolic-related regulation of inflammatory response in autoimmunity, such as RA and SLE-associated pathological processes. In many inflammatory conditions, the accumulation of ROS leads to oxidative stress. Generally, ROS is considered a pro-inflammatory mediator, which exacerbates inflammation via multiple parallel mechanisms. Firstly, ROS could promote the activation of inflammatory signaling pathways, such as TNF-induced NF-κB activation (Blaser et al., 2016). Secondly, ROS could actively mediate the oxidation of cellular contents, including neutrophilic mitochondrial DNA which makes a major contribution to type I IFN production in SLE pDCs (Caielli et al., 2016; Lood et al., 2016). Thirdly, dysfunctional mitochondria under oxidative stress are associated with reprogrammed metabolic profiles of immune cells in disease settings, which closely interact with local inflammatory environments and modulate the inflammatory phenotypes of immune cells (Huang and Perl, 2018). Given the accumulating evidence on metabolic pathway-regulated inflammation in autoimmune diseases, targeting reprogrammed metabolic profiles has been considered one of the promising therapeutic strategies for autoimmune disease treatment.

cistanche benefits for men-strengthen immune system
Glycolysis, oxidative phosphorylation, and ROS in RA and SLE T cells
Although glucose metabolism is one of the major energy sources during the immune response, the following metabolic processes are diverse, referring to glycolysis, oxidative phosphorylation, and pentose phosphate pathway. Glycolysis (or aerobic glycolysis) and oxidative phosphorylation produce ATP to fuel the immune response, but two pathways are polarized into two directions to govern the different inflammatory phenotypes of immune cells (O'Neill et al., 2016). In most cases, increased glycolysis is associated with inflammatory immune cells, and increased oxidative phosphorylation is associated with anti-inflammatory or non-inflammatory phenotypes. In T cells, the associations between metabolic status and T cell function in different subsets or different activation states have been extensively studied (Saravia et al., 2020). In short, glycolysis-featured Th1 and Th17 cells also require a glycolytic pathway for differentiation, and FOXP3 expression in Treg cells coordinates the programming of high mitochondrial oxidation and low glycolysis metabolism. In SLE patients, T cells exhibit elevated glucose metabolism compared with the T cells from healthy individuals (Doherty et al., 2014; Yin et al., 2015). Glucose uptake is mediated by glucose transporters, and the most well-studied glucose transporter, GLUT1, is highly expressed in SLE T cells (Koga et al., 2019). Interestingly, the mice over-expressing GLUT1 develop lupus-like phenotypes (Jacobs et al., 2008), such as autoantibody production and immune complex deposition in kidney tissues, which suggest the pathogenic role of enhanced glucose metabolism. Meanwhile, both the increased oxidative phosphorylation and increased glycolysis rate have been observed in SLE T cells (Doherty et al., 2014; Yin et al., 2015), but such increased glucose metabolism fails to generate ATP and leads to an ATP-depriving condition in SLE T cells, which is part of the prominent oxidative stress in SLE pathogenesis that mitochondria hyperpolarization accounts for failure of ATP generation and promotes production of ROS (Perl, 2013). Moreover, the accumulation of ROS could also result from an impaired ROS-neutralizing system, antioxidant-glutathione, and NADPH from the pentose phosphate pathway, in SLE patients (Gergely et al., 2002; Perl et al., 2015). Thus, increased glycolysis could promote Th1 or Th17 differentiation in SLE patients based on previous findings. Additionally, the oxidative stress manifested by the accumulation of ROS could also contribute to altered T-cell responses, where increased oxidative stress is associated with enhanced Th1/ Th17 response and SLE disease progression (Scavuzzi et al., 2018). The topic of the promotion of Th17 differentiation by ROS in various autoimmunity conditions has been previously summarized (Peng et al., 2021). Thus, with such upregulated glycolysis and redox signaling in SLE T cells, a Th1/Th17 dominant T cell response could be derived from these rewired metabolic pathways. T cells in RA patients also suffer the ATP shortage, which could result from defective mitochondrial biogenesis that is caused by deficiency of nuclease MRE11A in RA T cells (Li et al., 2016b; Li et al., 2019b). Another mechanism for ATP deprivation is rewired glucose metabolism in RA T cells. In contrast to enhanced glycolysis in SLE T cells, RA T cells show a low glycolytic metabolism profile (Yang et al., 2013; Yang et al., 2016). Actually, glucose metabolism is still utilized by RA T cells, but glycolysis is switched to the pentose phosphate pathway by insufficient expression of glycolytic enzyme 6-phosphofructo-2-kinase (PFKFB3) and upregulation of glucose-6-phosphate dehydrogenase (G6PD) (Yang et al., 2013). Consequently, hyperproduction of NADPH from the pentose phosphate pathway neutralizes ROS in RA T cells, and the redox-sensitive kinase, ataxia telangiectasia mutated (ATM), remains inactive during T cell proliferation, which allows the RA T cells to hyper-proliferate by bypassing the G2/M cell cycle checkpoint and further differentiate into Th1 and Th17 cells (Yang et al., 2016). Thus, glucose metabolism promotes RA pathogenesis via pentose phosphate pathway-mediated inflammatory T-cell response.
Metabolic abnormality in other immune cells in autoimmunity
In both RA and SLE-related disease conditions, besides T cells, the metabolic signatures of macrophages and other immune cells are largely unknown. Under inflammatory environments, activated macrophages adapt to the dysfunctional mitochondria and promote glycolysis to measure up to sufficient ATP production (Kelly and O'Neill, 2015). Meanwhile, the reduced rate of oxidative phosphorylation will make metabolites in the tricarboxylic acid (TCA) cycle accumulate, which could modulate inflammatory gene expression, such as succinate-stabilized HIF1α for Il1b expression (Murphy and O'Neill, 2018). Macrophages or monocytes could undergo similar metabolic reprogramming in autoimmune diseases. For example, in RA synovium, not only enriched lactate suggests a high level of glycolysis in tissues (Fujii et al., 2015; Haas et al., 2015; Kim et al., 2014), but the accumulation of intermediate metabolites in the TCA cycle is also evident in RA synovial fluids (Kim et al., 2014). Despite the to-be-determined macrophage-specific metabolic profile in RA synovium, autocrine or paracrine of succinate has been implied to mediate pathogenic IL-1β production in RA synovial macrophages via macrophage-expressed succinate receptor, GRP91 (Littlewood-Evans et al., 2016). Therefore, metabolic reprogramming in inflammatory tissues profoundly influences the pathogenic role of macrophages in autoimmune diseases, caused by the intrinsic or extrinsic metabolic signals (Liang et al., 2020). Further metabolic characterization of macrophages and other immune cells in autoimmune diseases would be informative to deepen our understanding of the immunome metabolism-related pathogenic mechanisms.
Future perspective: understanding autoimmunity at the single-cell level
Single-cell sequencing of mRNA expression levels for each individual cell in inflammatory conditions provides unique opportunities to dissect the inflammatory phenotypes in autoimmune diseases at an unprecedented resolution. Single cell expression profiles may aid in the identification of disease-specific cell subsets in lesion tissues and circulation as well as extrapolation of the complex intercellular communication networks. One of the successful examples of investigation of autoimmune disease pathogenesis with scRNA-seq is in RA synovial tissues. In 2019, the Accelerating Medicines Partnership Rheumatoid Arthritis and Systemic Lupus Erythematosus (AMP RA/SLE) Consortium published their multi-center investigations on the inflammatory characteristics of cellular components in RA synovial tissues (Zhang et al., 2019a). This study revealed the unexpected inflammatory phenotypes of local immune cells and fibroblasts, where HLA-DRAhi subliming fibroblasts, but not monocytes, are one of the major IL6 expressing cell types in RA synovial tissues, and CD8+ T cells, but not CD4+ T cells, are the major contributors for IFNG expressing cells in RA synovial tissues. Meanwhile, the expanded THY1 (CD90)+ HLA-DRAhi subliming fibroblasts, IL1B+ pro-inflammatory monocytes, ITGAX+ TBX21+ autoimmune-associated B cells, and PDCD1+ peripheral helper T cells and follicular helper T cells were identified by scRNA-seq as RA-associated cell subsets comparing to osteoarthritis patients' synovial tissues (Zhang et al., 2019a). The insightful findings on RA-associated cell subsets not only solidified the previously identified pathogenic role of peripheral helper T cells in RA synovium (Rao et al., 2017) but also guided the following studies pursuing the RA pathogenesis mediated by Notch-induced subliming fibroblasts and pathogenic HBEGF+ monocyte subset (Kuo et al., 2019; Wei et al., 2020). In this regard, the successful experience of scRNA-seq analysis for RA synovial tissues could be broadly applied to inflammatory tissues in autoimmune diseases, and advanced bioinformatics tools could be used to identify the prominent signaling pathways that are potentially involved in the formation and functionality of disease-associated cell subsets (Armingol et al., 2021). In the past few years, a subset of autoimmune diseases have been analyzed by the approach of single-cell expression profiling (Baglaenko et al., 2021). Studies of individual disease types have achieved certain success such as identification of disease-associated immune cell subsets. Nevertheless, illustrating generic versus disease-specific inflammatory phenotypes across various autoimmune diseases remains challenging given the difficulty of integrating distinct data sets from different sources (Stuart and Satija, 2019). Another promising direction for single-cell sequencing data analysis is single-cell immunome metabolism (Artyomov and Van den Bossche, 2020). In conventional metabolism profiling for immune cells, the metabolites are measured in bulk cell populations, which demands a large quantity of purified immune cells in certain types. However, the limited size and amount of tissue samples demand alternative experimental strategies to characterize the metabolic features of immune cells in inflammatory tissues from autoimmune disease patients. With the successful compiling of metabolic features from transcriptome data in immune cells, the current methodologies for transcriptome-based metabolic analysis are developed based on either pathway-based analyses or flux balance analysis (FBA)-based methods, which are conducted for evaluation of specific metabolic pathways or global characterization of interactively connected metabolic networks, respectively (Artyomov and Van den Bossche, 2020). Excitingly, both pathway-based analysis and flux balance analysis have been successfully applied for scRNA-seq-based investigations in disease-related immunome metabolism (Miragaia et al., 2019; Wagner et al., 2021). Therefore, combining single-cell expression profiles and advanced bioinformatics tools could be a powerful approach to dissecting the metabolic status of immune cells in autoimmune diseases.
Genetic risk factors have been identified in various autoimmune diseases, among which some gene loci encode components of multiple pathways with specified functionality in the immune system. The abnormalities in these pathways are associated with a break of self-tolerance and excessive inflammation, contributing to the pathogenesis, such as HLA alleles and ERAP1/2-related antigen presentation, and IFNAR1-related type I IFN pathway. Meanwhile, some genetic variants could not be directly aligned to autoimmunity and inflammation. The pathways behind these variants are being functionally associated with autoimmune diseases with accumulating knowledge, such as Notch signaling and metabolic pathway-related immuno-pathogenesis. In the future, annotating the variants with unknown functionality in the pathological processes will provide an in-depth dissection of predisposed disease-prone conditions with therapeutic insights.






