Parasites, Infections, And Inoculation in Synthetic Minimal Cells
Oct 27, 2023
ABSTRACT: Synthetic minimal cells provide a controllable and engineerable model for biological processes. While much simpler than any live natural cell, synthetic cells offer a chassis for investigating the chemical foundations of key biological processes. Herein, we show a synthetic cell system with host cells, interacting with parasites and undergoing infections of varying severity. We demonstrate how the host can be engineered to resist infection, we investigate the metabolic cost of carrying resistance, and we show an inoculation that immunizes the host against pathogens. Our work expands the synthetic cell engineering toolbox by demonstrating host−pathogen interactions and mechanisms for acquiring immunity. This brings synthetic cell systems one step closer to providing a comprehensive model of complex, natural life.

cistanche benefits for men-strengthen immune system
INTRODUCTION
Within the past several decades, the field of synthetic biology has exploded, allowing for the construction of synthetic cells that resemble and model aspects of natural biological life. Modeling cellular pathways and behaviors in synthetic cells offers many advantages over live cells such as a more simplified and defined reaction environment, complete control over the proteomic and chemical makeup of the cell, the ability to compartmentalize interfering biological pathways and processes, as well as maintaining important biological systems that are absent in in vitro reaction systems.1,2 These unique assets of synthetic cell technologies have already allowed for simulation of biological conditions and pathways within a life-like in vitro bioreactor (synthetic cell) and brought a better understanding to many areas of study, including molecular crowding,3 lipid−protein dynamics,4 minimal metabolism,5 and many other critical aspects of biological life.2 These life-like synthetic cells with their unique assets are rapidly becoming better models and shedding more light into the mysterious inner workings of cellular life.6,7 These breakthroughs are not confined to single-cell processes; engineering interactions between populations of liposomes have proved a powerful technology as well. For example, the design of a synthetic cellular system that mimics a predator−prey relationship has been developed utilizing light as the signal between these synthetic cell populations.8 Another complex work has been developed that allows for signal propagation throughout a synthetic cell droplet matrix, allowing fluorescence to spread in a spatial manner among the population.9 In addition to synthetic cell−synthetic cell interactions, there have been recent advancements in synthetic cell−live cell interfaces as well. For example, it has been shown that a synthetic cell designed to interface with a neural stem cell is able to induce neural differentiation as well as act as a generalized chassis capable of interfacing with other cell types.10 There have been countless advancements in the development of signal propagation and synthetic cells acting as interfaces, much of which is explored in the following reviews.11−13 In short, there are whole fields of research concerned with mimicking the interactions of cellular populations in the model system of synthetic cells. Despite this, there has not been a study that uses synthetic cells to model infection, parasitism, or inoculation. Just as synthetic cells are commonly used to model complex biological processes in a simplified and controlled way, the potential exists to be able to model these complex interorganism interactions. The importance of this can be seen in the groundbreaking study that successfully produced active phage particles within a cell-free protein expression platform.14 We aim to bring this concept to the macro level by designing synthetic cell populations that can actively infect one another and hijack a host cell’s genome, just as live viruses and parasites do. Being able to apply and engineer synthetic cells to model these disease behaviors will provide a valuable tool to the scientific community.
In this work, we exhibit our system for synthetic cell disease modeling. We engineered a host cell that could undergo infection from the second population of parasitic synthetic cells and investigated the resource competition, genome hijacking, and metabolic repurposing that occurs during such an infection. We also model a method that allows for the rescue of the host after infection as well as a means of inoculation that will allow for the host to resist the infection of the disease-causing synthetic cells. In short, we present a novel disease and infection model chassis for synthetic cells. With this technology, synthetic cells can begin to be used as rudimentary models for disease, infection, and inoculation (Figure 1).

Figure 1. Parasite, infection, and inoculation system in synthetic minimal cells. (a) The “host” is a synthetic cell expressing green fluorescent protein (GFP). On the surface of the outer membrane, the host carries ssDNA tags. The only proteins translated into a healthy host are the host’s own proteins from the plasmids the host was created with. (b) The “parasite” is a liposome filled with plasmid DNA encoding a gene for mCherry fluorescent protein, and the surface of the parasite liposome is decorated with ssDNA tag (complementary to the tag on the host surface). The “infection” starts with a fusion of the host and parasite liposomes, mediated by the complementary DNA tags on their surface. After the infection, the host expresses both the original host gene (GFP) and the parasite gene (mCherry). (c) The “infection” starts with “pathogen” liposomes carrying a gene for the restriction enzyme MunI fusing with the host liposomes. The pathogen gene MunI is expressed by the host and the MunI restriction enzyme digests host DNA. This results in the native host genome being digested, turning most of the host protein expression from GFP to the pathogen’s encoded protein, MunI. (d) The “lethal infection” is a fusion of the host liposome (with the GFP genome) with the pathogen liposome containing the gene encoding aHL, a membrane protein. After the infection, the host expresses aHL, which creates membrane pores, causing leakage of host nutrients and effectively shutting down the host’s metabolism. (e) “Immunization” is the incubation of the host with ssDNA containing a sequence compatible with the fusion tags on the surface of the host liposomes. The immunizing DNA creates a duplex with host DNA tags, preventing the hybridization of other DNA oligos to these tags. This prevents the fusion of any pathogenic liposomes to the host. This is a proof-of-concept figure, and the graphs are for illustrative purposes, representing relative amounts of protein expressed from the host and various pathogens. Real data for each system are provided in subsequent figures.
RESULTS AND DISCUSSION
In this work, we use the words: host, parasite, infection, deadly, vaccine, and inoculation all in the context of nonliving, synthetic cell systems. By “deadly” infection, we mean a marked decrease in translation in the population of synthetic cells. All experiments in this work were performed on synthetic cells that are not alive, do not self-replicate, and do not evolve. This is a model system for natural biological properties and processes.

cistanche supplement benefits-increase immunity
Click here to view Cistanche Enhance Immunity products
【Ask for more】 Email:cindy.xue@wecistanche.com / Whats App: 0086 18599088692 / Wechat: 18599088692
Infection of a Synthetic Cell with a Parasitic Synthetic Cell Hijacks Host Resources.
To test the concept of a synthetic cell system mimicking an infection, we started by constructing a model system for a parasite. We used the simplest concept of a parasite possible to reconstitute in this nonliving biochemical system: a parasite that hijacks some of the resources of the host. In this case, the host metabolism is represented by the expression of GFP protein, and the parasite is represented by mCherry protein expression. For this system, we utilized the previously demonstrated phenomenon of linear protein expression yield dependent on the DNA template concentration in TxTl.15,16 First, we set out to find the optimal total plasmid concentration at which the ratio of GFP and mCherry DNA corresponds to the ratio of observed fluorescence from these proteins. We arbitrarily assigned the GFP gene as the “host” and the mCherry gene as the “parasite”. These names signify nothing special in this particular experiment but will indicate the host and parasite in subsequent synthetic cell experiments. Here, we use these names to make it easier to relate these proof-of concept experiments to subsequent synthetic cell infection experiments using the same plasmids. We mixed GFP and mCherry plasmids, with both genes under the control of the T7 promoter at varying plasmid ratios and a total plasmid concentration in the samples 1, 5, 10, or 15 nM (Figure 2b). The results followed the expected nearly linear relationship between the concentration of plasmid and the recorded fluorescence intensity, with the best match between fluorescence and plasmid concentration ratios observed at a total of 10 nM plasmid. The observed relationship indicates that the plasmids, with the proteins being similar in size and amino acid composition (Figure S1), compete for the same pool of TxTl resources for their expression. Changing the relative concentration of the two plasmids results in changes in the expression efficiency roughly proportional to the ratio of the plasmids. This indicates that it would be a good model to mimic parasite infection: the parasite genome (mCherry plasmid) will compete with the host genome (GFP plasmid). Next, we decided to test a case where either of the plasmids was under the control of a stronger promoter, thus mimicking a biased resource allocation scenario. To keep the resource competition aspect of the future “parasite” infection experiments, we kept both genes under the same RNA polymerase promoter (the T7 RNA polymerase, T7 RNAP) but varied its strength. We used a strong T7 RNAP promoter T7Max, which demonstrates significantly higher transcription yield, resulting in correspondingly higher translation yields, in TxTl.17 We tested the reaction setup identical to the one described earlier: two plasmids at different ratios, tested at varying final total plasmid concentrations. One of the plasmids was under the control of the strong T7Max promoter, while the other was under the canonical T7 promoter. In both test cases, the T7Max plasmid expression was strong enough that the fluorescence of the protein from a gene under T7Max did not significantly increase with the increased total concentration of the plasmid (Figure 2c,2d). It did not seem to matter if T7Max was driving GFP or mCherry, both proteins responded to the stronger promoter with a similar relative increase. This provided a promising model for infection with a “stronger” parasite (a parasite carrying gene under T7Max into a host under T7) and reverse, stronger host resisting weaker parasite (a host with genome under T7Max and a parasite carrying T7 gene). Encouraged by these proof-of-principle experiments, we moved on to synthetic cell experiments.
The “infection,” in all cases, is the fusion of a host and parasite synthetic cell. The fusion is facilitated by DNA tags on the surface of the liposomes, using the DNA-inducible fusion system described before.18 Briefly, the synthetic cells were created with Escherichia coli TxTl inside a membrane of POPC and cholesterol, and the outer leaflet was decorated with single stranded DNA anchored to the membrane by cholesterol modification. If a pair of liposomes with complementary tags meet, the DNA tags hybridize, bringing the membranes together and initiating membrane fusion, which is followed by lumen mixing. In our experiments, the host synthetic cell is the liposome containing TxTl with T7 RNAP in addition to the genome of a plasmid-encoded GFP. Monitoring GFP fluorescence serves as a proxy for the “health” of the host metabolism. The incoming parasite contains a genome in the form of a plasmid encoding mCherry, the complete translation system, but no T7 RNA polymerase. This ensures that any observed changes in the host metabolism result from the genes brought in by the parasite and not from the fusion of the parasite and host liposomes, diluting the host cytoplasm. The parasite infection experiments were carried out with the host membrane labeled with Marina Blue DHPE, a blue membrane lipid dye. This allowed us to normalize the observed host and parasite gene fluorescence to the blue fluorescence, normalizing the results to the total amount of host membrane fluorescence present in the sample. This accounts for the dilutions of the total volume of the sample after the addition of parasite liposomes. The parasite membranes were not fluorescently labeled. All experiments were accompanied by a control sample, where only one population was present the host, but with mCherry plasmid, to show the theoretical maximum possible mCherry expression levels if it was the only gene in the population. In the first set of experiments, we mixed the host and parasite with genes both under the control of the regular T7 promoter so both host and parasite genes were equally “strong.” We mixed the host with the parasite so that the parasite was at 25, 50, or 75% of the total liposome population. In all cases, the host GFP fluorescence decreased in the presence of the parasite, with the corresponding increase of parasite mCherry fluorescence (Figure 2e). Samples with only the parasite and no host produced no fluorescence of either color, as expected. In all cases, the DNA-mediated liposome fusion is not 100% efficient;18,19 Therefore, the relative decrease in host fluorescence and increase in parasite fluorescence were not as large as the expected theoretical value based on the ratio of host to parasite. In other words, the infection was not 100% efficient. This serendipitous quality of the DNA-induced fusion system was, in this case, a beneficial feature of the system. As in the case of natural infection, the synthetic cell fusion-mediated infection did not affect all hosts and not all parasites found a host. Next, we investigated two cases, where the host and parasite were not evenly matched in strength of their genomes. In one case, the host genome was under the control of the T7 promoter while the parasite genome was under the T7Max promoter (Figure 2f), and in the opposite scenario, the host genome was under T7Max and the parasite genome was under the T7 promoter (Figure 2g). In the case of the weaker host attacked by a stronger parasite, the relative increase in parasitic mCherry fluorescence and the corresponding decrease in host GFP fluorescence were much stronger than in the previous case of an evenly matched host and parasite. As expected, when the host was stronger than the parasite, the opposite was true: the mCherry fluorescence levels were lower and the relative decrease in GFP fluorescence was lower than in the evenly matched case. This is best illustrated by comparing a single data point under the same infection conditions: at 75% parasite ratio. Evenly matched host and parasite populations result in a parasite fluorescence slightly larger than the host. When the parasite is stronger, the parasite fluorescence is significantly higher than the host. And when the host is stronger, the parasite fluorescence is slightly lower than the host. These highlighted data points are marked with a yellow star in Figure 2e−g. We performed the control experiment of a host infected with an “asymptomatic” parasite a parasite carrying a plasmid with the T7 promoter but no protein coding sequence (we removed the mCherry cassette from the plasmid) (Figure 2h). The host GFP fluorescence decreases minimally at higher parasite levels, indicating that some small amount of host resources are devoted to transcribing the short noncoding RNA from the parasite plasmid. This data also confirms that the main metabolic burden of the parasite infection on the host lies in utilizing translational, not transcriptional, resources. This observation falls in line with previous observations of translation being the more variable and limiting resource in cell-free systems.20 Overall, these experiments provide a model for infection with a parasite that hijacks the metabolic resources of the host, with synthetic cell behaviors mimicking aspects of a natural host-parasite relationship.

Figure 2. Parasite infection hijacks the resources of the host. (a) The host synthetic cell liposome contains a cell-free translation system and a plasmid encoding GFP. The surface of the host is decorated with ssDNA tags. The parasite liposome contains plasmid encoding mCherry, with a translation system but no RNA polymerase (so the parasite cannot express mCherry). The surface of the parasite is decorated with ssDNA tags complementary to the tags on the surface of the host. The DNA tags facilitate the fusion of the host and parasite liposomes and the host expresses both its own genomic GFP and the parasitic genome containing mCherry. (b−d) Proof-of-principle for the host (GFP) and parasite (mCherry) genomes competing for the same synthetic cell resources. These experiments are done in solution TxTl, not in liposomes. Two plasmids, encoding GFP and mCherry, were mixed at the ratio indicated by the plasmid icons on the left of the heat map, with the sample’s total DNA concentration indicated in the row above the heat map. Fluorescence was measured in green (GFP) and red (mCherry) channels. Each plasmid was tested in two variants with promoters of different strengths: with T7 and with the stronger T7Max promoter. Individual data for all heat maps are provided in Figures S2−S4. Representative individual time courses for the protein expression of the T7 host and parasite are provided in Figures S5−S7 and for the T7Max parasite in Figures S8−S10. (e−g) Infection of the host with the parasite. In each experiment, host liposomes are mixed with parasite liposomes. The fusion of the parasite to the host delivers parasitic mCherry DNA, which is expressed alongside host GFP DNA. Different ratios of the host to the parasite were tested, as indicated by the icons below the bar graphs. The host genome under the T7 promoter was infected with a parasite with its genome under the T7 promoter panel (e) and with a parasite with its genome under the T7Max promoter panel (f), the host was also made with a T7Max genome and infected with the parasite with a T7 genome panel (g). Representative individual time courses for protein expression are provided in Figure S11. Similar experiments at higher and lower total lipid concentrations are provided in Figure S12. Size-exclusion purification traces showing vesicle stability postfusion are provided in Figure S13. (h) Infection with incompatible parasite: the parasite carries the T7 promoter but noncoding DNA. Error bars indicate SEM, n = 3. Fluorescence values in panels (e−h) are normalized to the blue fluorescence of a membrane dye to normalize the results to the concentration of host cells. All sample series labeled “host GFP infected” are experiments, where the host and parasite had complementary DNA tags on the surface, enabling fusion of the liposomes, termed infection. Series-labeled “host GFP healthy” show samples, where the host and parasite were mixed at the ratios shown, but did not contain complementary fusion tags, making infection impossible. The diamond in panels (e−h) indicates one particular infection condition that best illustrates the differences in the host and parasite expression strengths in panels (e−g) and the benefits of host immunity in panel (h). A direct comparison of the host and parasite values signified by the star is given in Figure S14. Data for infection with a parasite without any plasmid are shown in Figure S15.
Infection of a Synthetic Cell Can Destroy the Host Genome.
The parasite described above was expressing a genome-encoding mCherry using resources of the host, but the host maintained some of its metabolism and continued expressing GFP despite the infection. Next, we decided to investigate a system in which the infection results in a complete takeover of the host metabolism by the parasite via the destruction of the host genome (Figure 3a). To engineer such a scenario, we investigated a restriction enzyme digest of a plasmid in TxTl. In the proof-of-principle experiments, a MunI restriction enzyme was expressed in TxTl without liposomes. The MunI plasmid was mixed with GFP plasmid at different ratios. If the GFP plasmid contains no MunI restriction sites, the relatively small decrease in observed GFP fluorescence can be accounted for by the previously described competition between the two plasmids for TxTl resources. If the GFP plasmid has a MunI restriction site, the GFP fluorescence decreases significantly, and the decrease scaled proportionally to an increasing amount of the MunI-encoding plasmid (Figure 3b). The MunI expression was monitored via Western Blot and scaled proportionally with the plasmid concentration (Figure 3c).

cistanche tubulosa-improve immune system
Encouraged by these results, we set up an infection experiment with synthetic cells. The host carried the GFP plasmid and the pathogen carried the MunI plasmid. The concentration of GFP and MunI plasmids was set to 2 mM, well below the concentration of plasmid saturation for TxTl.21 We intended to remain under the total plasmid concentration at which host and parasite expression would be high enough to directly compete for limited resources (like in the above-described case of a simple parasite), instead intending to mostly measure the effect of the parasite enzyme actively destroying the host genome. Like in the earlier described parasite experiments, the host contained bacterial TxTl and T7 RNA polymerase, the pathogen carried only cell extract without T7 RNA polymerase, and both the host and pathogen were labeled with complementary DNA fusion tags. The host membrane was labeled with Marina Blue DHPE to normalize fluorescence results to the concentration of host liposomes. The host and pathogen were mixed at varying ratios, with the pathogen being 25, 50, or 75% of the total population. The fluorescence of the “infected” host, where the host and pathogen contained complementary fusion tags, decreased with an increasing amount of pathogen. The GFP fluorescence from the “healthy” host, where the pathogen did not contain complementary fusion tags, did not decrease in the presence of any amount of pathogen. Samples with only the pathogen resulted in no measurable fluorescence (Figure 3d). The pathogenic MunI protein expression was quantified using Western Blot analysis, with data shown for the sample series of the infected host (Figure 3e). Next, we investigated infection in the case of a host immune to the pathogen: the host GFP plasmid did not have a MunI restriction site (Figure 3f). The host fluorescence after infection decreased but was nowhere near as significant as in the earlier case of a host with a MunI site in the genome. The MunI pathogen protein was expressed at levels comparable to earlier infection experiments (Figure 3g), so the metabolic load on the host was similar. The infected host fluorescence decrease in these experiments is caused by some of the host resources being hijacked by the pathogen expressing MunI, but because the total plasmid concentration is much lower than in the earlier simple pathogen experiments, the host fluorescence decrease is less significant. This experiment also confirms that the decrease in fluorescence of the host with a MunI site (Figure 3d) was caused by protein expression time courses provided in Figure S22. (g) Expression of MunI was tracked via western blot analysis through the experiment for the samples shown in the series “GFP infected” in panel (f). Error bars on all panels indicate SEM, n = 3. Fluorescence values in panels (d, f) are normalized to the blue fluorescence of the membrane dye used to normalize the results to the concentration of the host cells. Fluorescence in panel (b) is normalized to 0 mM MunI plasmid for each sample series. The diamond symbols in panels (d, f) highlight one particular condition best illustrating the differences in responses to the infection from a host susceptible to the pathogen panel (d) and immune to the pathogen panel (f). All of the sample series labeled “GFP infected” are experiments, where the host and pathogen had complementary DNA tags on the surface, enabling the fusion of the liposomes, termed infection. The series labeled “GFP healthy” shows samples, where the host and pathogen were mixed at the ratios shown but did not contain complementary fusion tags, making infection impossible.

Figure 3. Infection destroys the host genome and hijacks host resources. (a) The host cells contain plasmid encoding GFP, with two MunI restriction sites; the pathogen contains plasmid encoding the MunI restriction enzyme. After the fusion of the host and pathogen, the host TxTl expresses the MunI restriction enzyme, which then digests the host genome. (b) The proof-of-principle experiments for the infection: GFP plasmid was prepared with and without MunI restriction sites. GFP plasmid was mixed with MunI plasmid and GFP expression was recorded. These experiments were performed in solution TxTl without liposomes. Proof-of-principle for the infection experiment, with the MunI enzyme added externally instead of being expressed in TxTl, is shown in Figures S16 and S17. Representative protein expression time courses are provided in Figure S18. (c) MunI expression was monitored by western blot analysis; the intensity of the MunI band was quantified in the same samples as fluorescence results shown in panel (b) for the “MunI site absent” series. A representative western blot of the MunI restriction enzyme is provided in Figure S19. (d) GFP fluorescence of the host mixed with pathogen liposomes at the ratio shown under the data graph. The host GFP plasmid has two MunI restriction sites. Representative protein expression time courses are provided in Figure S20. Data for infection experiments with incompatible fusion tags are provided in Figure S21. (e) Expression of MunI was tracked via western blot analysis through the experiment for the samples shown in the series“GFP infected” in panel (d). (f) GFP fluorescence of the host mixed with pathogen liposomes at the ratio shown under the data graph. The host GFP has no MunI restriction sites, giving it immunity to the pathogen protein. Representative protein expression time courses are provided in Figure S22. (g) Expression of MunI was tracked via western blot analysis through the experiment for the samples shown in the series “GFP infected” in panel (f). Error bars on all panels indicate SEM, n = 3. Fluorescence values in panels (d, f) are normalized to the blue fluorescence of the membrane dye used to normalize the results to the concentration of the host cells. Fluorescence in panel (b) is normalized to 0 mM MunI plasmid for each sample series. The diamond symbols in panels (d, f) highlight one particular condition best illustrating the differences in responses to the infection from a host susceptible to the pathogen panel (d) and immune to the pathogen panel (f). All of the sample series labeled “GFP infected” are experiments, where the host and pathogen had complementary DNA tags on the surface, enabling the fusion of the liposomes, termed infection. The series labeled “GFP healthy” shows samples, where the host and pathogen were mixed at the ratios shown but did not contain complementary fusion tags, making infection impossible.
Infection of a Synthetic Cell Can be “Deadly”, Depleting Host Cell Resources, and Such Infections Can be “Treated” to Rescue Function.
The above parasite and pathogen experiments demonstrated that synthetic cells can be used as a model for infection, where a pathogen hijacks the resources of the host. Next, we decided to investigate a scenario in which the infection causes the metabolism of the host to cease functioning after the expression of the pathogenic protein. In this case, the pathogen is comparable to infection with a bacterium expressing a deadly toxin minus the replication aspect of the host or parasite since these synthetic cells do not replicate (Figure 4a). To engineer this system, we chose α hemolysin (aHL), a natural toxin and membrane protein widely used in synthetic cell engineering to create nonspecific membrane pores.22,23 We appreciate the irony of this choice as aHL is naturally expressed by Staphylococcus aureus as a toxin, and the protein was later appropriated to engineer membrane transport and prolong synthetic cell metabolism.24 Here, we revert to using aHL for its natural role as a harmful toxin produced by an infection. Another notable synthetic cell engineering-related example of using aHL’s toxicity to cause natural cell death comes from the application of synthetic cells as a cancer therapy technology.25 As a proof of principle, we established that expressing a high concentration of aHL in synthetic cell vesicles results in leakage of cell content and a significant decrease in the expression of GFP inside the cell. This effect can be reversed if the outside of the liposomes contains all of the small molecules necessary for TxTl (Figure 4b). In our host−pathogen model system, the host contains a GFP encoding plasmid and the pathogen contains an aHL-encoding plasmid. All of the infection experiments’ design is similar to the parasite and pathogen scenarios described above, with the pathogen carrying no T7 RNAP and both the host and pathogen decorated with DNA fusion tags. Interestingly, we observed that at lower pathogen concentrations (the pathogen being 25 or 50% of the total population), the host fluorescence decreases only slightly. However, increasing the pathogen concentration to 75% results in a significant decrease in the host GFP fluorescence (Figure 4c). We speculate that this result might be caused by the relatively slow leakage of nutrients from synthetic cells at a lower aHL concentration. The GFP fluorescence was measured only after 12 h of incubation and GFP is a very stable protein. Therefore, it is possible that in the case of lower aHL concentrations (lower pathogen amount), the aHL-caused leakage of nutrients was slow enough that the host cells accumulated significant amounts of GFP before the effects of the aHL-induced leakage became crippling to the metabolism. To decouple the results of simple competition for resources between the host and parasite genomes, we performed “infection with rescue” experiments in which the outside buffer contained all of the small molecules that compose the TxTl energy, amino acid, and salt mixtures. This was set up similarly to the “nutrients outside” condition tested in Figure 4b. The decrease in host fluorescence in these experiments was similar to the earlier “infection without rescue” samples with 25 and 50% levels of the pathogen. The decrease of host fluorescence at 75% of the pathogen was significantly smaller than the decrease observed in samples without rescue (Figure 4d). The big difference in results for the experiment with 75% of the parasite, with data points in Figure 4 panels (c, d) highlighted with a star, seems to support our hypothesis that higher aHL concentrations cause harm early in the reaction. At a lower pathogen concentration, the leakage is slow enough that the host GFP accumulates (as seen both with and without rescue nutrients), so the observed GFP fluorescence decreases after infection mostly due to the competition for resources. At higher pathogen concentrations, the GFP fluorescence in samples with external nutrients decreases mostly due to resource competition. These experiments demonstrate that we can engineer synthetic cells to model infection, resulting in two different outcomes: the metabolic penalty of the infection itself (decreased GFP due to resource competition) and a more “deadly” infection resulting from abolishing the hosts’ protein expression. Encouraged by this observation, we decided to investigate a system in which synthetic cells could actively resist a pathogenic infection and investigate the metabolic penalty of such a response.

cistanche benefits for men-strengthen immune system
Synthetic Cells Can Defend against Infection Using a Bacteria-Inspired Strategy.
Inspired by the well-known function of restriction endonucleases to safeguard bacteria against foreign DNA, we sought to employ this strategy within synthetic cells. We expanded on the parasite system presented in Figure 2: in addition to GFP, we added MunI-containing plasmid to the host. The parasite contained mCherry, as in previous experiments. The mCherry plasmid contains a MunI restriction site (Figure 5a). First, we performed proof-of-principle experiments to assess the metabolic load of expressing MunI on the host. In these unencapsulated TxTl experiments, we monitored the expression of GFP and expression of mCherry, each mixed with an increasing amount of either MunI or EcorI plasmid. All genes were under the control of the same T7 promoter. The GFP gene had an EcorI but not a MunI restriction site and the mCherry gene had a MunI but not an EcorI restriction site. When the restriction enzyme plasmid was mixed with a fluorescent protein plasmid without a recognition site for that restriction enzyme, the fluorescent protein expression decreased proportionally to the increase of restriction enzyme plasmid, indicating competition between the two plasmids for TxTl resources, as demonstrated earlier for GFP and mCherry in Figure 2. These noncutting pairs were GFP with MunI (Figure 5b) and mCherry with EcorI (Figure 5e). When a fluorescent protein was paired with a restriction enzyme capable of cutting the gene for that fluorescent protein, the fluorescence decreased significantly to nearly background levels at higher restriction enzyme plasmid concentrations. These cutting pairs were GFP with EcorI (Figure 5c) and mCherry with MunI (Figure 5d). This indicated a decrease in fluorescence significantly beyond the simple competition for resources and similar to the infection scenario demonstrated in Figure 3. MunI expression in the presence of GFP decreases GFP expression. If we use GFP as a proxy for host “baseline” metabolism, then MunI can be seen as a metabolic load on the host.

Figure 4. The host is susceptible to “deadly” infection, but the host can be rescued. (a) The host contains GFP plasmid, and the pathogen contains plasmid encoding the α hemolysin (aHL) membrane channel. After the infection, the host expresses aHL, which creates pores in the membrane, causing leakage of nutrients and resulting in a decrease in metabolic activity of the host. (b) Proof of principle for a host-expressing aHL (no infection, host was created with both 5nM GFP and the indicated amount of aHL plasmid). The host was incubated in a buffer with or without nutrients (all components of the energy, salt, and amino acid mix used to prepare the host cell-free translation system). The optimum rescue nutrient concentration was established experimentally, data shown in Figure S23. (c) Infection of the host with the pathogen, with varying amounts of pathogen per host cell. The leakage of small molecules from the “healthy” and “infected” cells was directly measured using a small molecule dye, data are shown in Figures S24 and S25. (d) Experiments similar to these are shown in panel (c), but the outside buffer contains small molecule nutrients similar to “nutrients outside” experimental conditions shown in panel (b). Error bars on all panels indicate SEM, n = 3. The fluorescence values in panels (b−d) are normalized to the blue fluorescence of the membrane dye used to normalize the signal to the concentration of the host cells. The diamond symbols in panels (c, d) highlight one particular condition best illustrating the differences in response to the infection from a host without outside nutrients panel (c) and immune to the pathogen due to the presence of outside nutrients panel (d). For panels (c, d), the series “GFP infected” is the host fused with the pathogen carrying aHL protein, and “GFP healthy” is the host fused with the pathogen carrying no plasmid.
We proceed to the infection experiments, mixing the host (with GFP and MunI plasmids) with a parasite (with mCherry plasmid). As previously, we investigated scenarios with an increasing amount of parasites in the population (Figure 5f). The positive control of a host-expressing mCherry only (with EcorI instead of MunI to balance the metabolic load) created a benchmark for the theoretical maximum amount of pathogen protein that can be expressed in the host (Figure 5f). As the amount of parasite increased, the host’s GFP fluorescence decreased, but the decrease in infected host fluorescence was significantly smaller than the decrease of GFP expression observed in earlier experiments with the same parasite. GFP and parasite mCherry fluorescence of this infected host-expressing MunI (data point labeled with a star in Figure 5f) can be directly compared with similar infection hosts without MunI (data point labeled with a star in Figure 2e). In the host expressing MunI, the host GFP started with an absolutely lower level while healthy (because some resources are diverted to expressing MunI) but after the infection, the host fluorescence decrease is much less significant, and the host makes much less parasite mCherry protein than in the case of the host not expressing MunI. This experiment demonstrates the benefit of the host spending some resources on defense against parasites. This system could be seen as a very simple biochemical model for organisms expressing proteins to defend against future infections or even harmful environmental factors. The host originally is less robust (if we take GFP expression as a benchmark for host health), but in case of an infection, the additional resources the host spent on expressing defensive proteins pay off, providing protection against the infecting parasite. Therefore, in this simple nonliving synthetic cell, we reconstituted an example of a fundamental biological arms race.

Figure 5. Host cells defend against infection. (a) The host cells contain two plasmids: one encoding GFP (without a MunI restriction site) and the other encoding the MunI restriction enzyme (the “parasite resistance” gene). The parasite contains mCherry plasmid, in a system similar to experiments shown in Figure 2. The infection is mediated by fusion via DNA tags on the surface of the host and parasite. After the infection, the mCherry plasmid introduced by the parasite is digested by the host’s MunI restriction enzyme. (b−e) Measuring the metabolic cost of expressing the parasite resistance. Experiments are in a bulk TxTl system, not encapsulated in vesicles. In each experiment, the fluorescent protein plasmid (GFP or mCherry) was mixed with restriction enzyme plasmid (MunI or EcorI) at the ratio indicated on the x-axis for the total plasmid concentration of 10 mM. GFP has an EcorI restriction site but not a MunI site, and mCherry has a MunI but not an EcorI restriction site. The trendline is a visual guide but not a data fit. Error bars indicate SEM, n = 3. Control experiments with purified restriction enzymes, instead of expressing them from a plasmid, are provided in Figure S26. The stability of liposomes with an increased amount of parasite plasmid was also measured, data shown in Figure S27. (f) Infection of the host with an increasing amount of parasite. “Host GFP healthy” is the expression of GFP from host cells fused with the parasite without any plasmid. “Host GFP infected” is the host fused with the parasite carrying mCherry. The red fluorescence is mCherry fluorescence from the parasite fused with the host (only data from the infected host are available. The healthy host samples did not carry mCherry plasmid). Error bars on all panels indicate SEM, n = 3. Fluorescence values are normalized to the blue fluorescence of the membrane dye used to normalize the results to the concentration of the host cells. The diamond symbol indicates the expression of the parasite protein under particular infection conditions, directly comparable to the data point highlighted with a star in Figure 2e an experiment under the same conditions, but with the host lacking the MunI resistance gene.
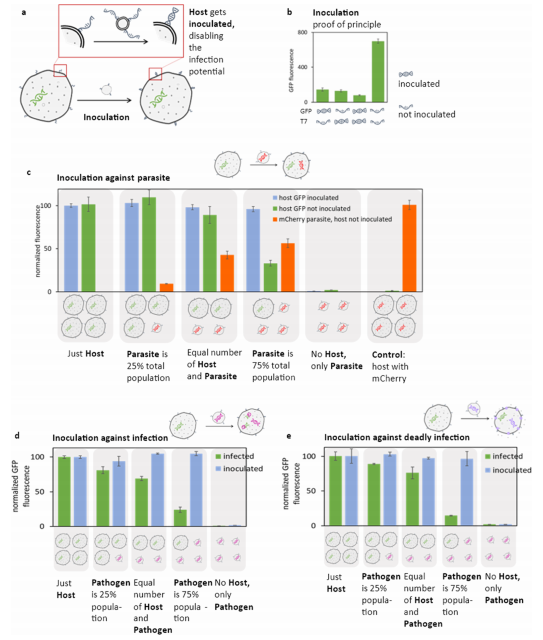
Figure 6. Inoculation provides defense against infections. (a) The host contains GFP plasmid. The pathogen in these experiments is one of the three infection schemes demonstrated in this paper: a parasite (as shown in Figure 2), a pathogen with the MunI restriction enzyme digesting the host’s GFP genome (as shown in Figure 3), or a pathogen carrying the membrane channel aHL causing leakage of nutrients from the host (as shown in Figure 4). In each case, the host and pathogen contain complementary DNA fusion tags. Before introducing the pathogen, the host is “inoculated”: incubated with excess ssDNA oligos complementary to the fusion DNA tags on the surface of the host (tags identical to these on the pathogen minus the cholesterol moiety used to anchor the tag to the membrane of the pathogen). The inoculation causes the fusion tags on the surface of the host to become double-stranded, thus making the fusion with the pathogen impossible. (b) The proof-of-principle for inoculation scheme. The population of liposomes carrying GFP under the control of the T7 RNA promoter, but without T7 RNA polymerase, is fused with liposomes containing T7 RNA polymerase. The GFP protein can be expressed only if the two populations fuse. One or both of the mixing partners were “inoculated” by incubation with a DNA tag compatible with the fusion tags on the surface of the liposome, rendering the fusion tags double-stranded and thus incompatible with fusion. A representative time course for protein expression is shown in Figure S28. Control experiments without any fusion tags are in Figure S29, and controls with incompatible fusion tags are provided in Figure S30. The inoculation oligo concentration was established experimentally, data shown in Figure S31. (c) Parasite infection experiments. A host carrying GFP was fused with a parasite carrying mCherry, as described in Figure 2. The host and parasite were mixed with a varying ratio of the host to the parasite; host cells were either inoculated (with double-stranded DNA fusion tags not compatible with fusion) or not inoculated (thus susceptible to fusion with the parasite). mCherry fluorescence is reported only for samples with the host not inoculated because there was no measurable mCherry fluorescence in the inoculated host samples. A reversed inoculation was tested by inoculating the parasite instead of the host, data shown in Figure S32. (d) Infection experiments with the host genome susceptible to the pathogen carrying the MunI restriction enzyme, as described in Figure 3. The infected data series represents a host without inoculation, and the inoculated data series represents samples with the host incubated with inoculation oligos, thus not susceptible to fusion with the pathogen. (e) Infection experiments with the pathogen carrying aHL gene, result in aHL channels leaking nutrients out of the host cells after infection, as described in Figure 4. The infected data series represents the host without inoculation and the inoculated data series represents samples with the host incubated with inoculation oligos, thus not susceptible to fusion with the pathogen. Error bars on all panels indicate SEM, n = 3. Fluorescence values in panels (c−e) are normalized to the blue fluorescence of the membrane dye used to normalize the results to the concentration of the host cells. Some data shown in this figure are experiments with setup identical to the experiments shown in earlier figures (as indicated in each caption). These experiments were repeated for this figure to produce data using the same batch of TxTl for directly comparing the results, eliminating batch-to-batch variability of TxTl preparations.
Inoculation of Synthetic Cells Prevents Infection.
In the above-described example, we demonstrated the host defending against the influence of a pathogen’s genome by expressing an extra protein. Next, we asked if it is possible to model a scenario, where some intervention makes a previously susceptible host immune to infection in the first place. We called it an inoculation model. While recognizing this is far from a perfect analogy, the aspect we focused on is that the host synthetic cell does not expend any resources on generating immunity. In our immunization model, we designed a treatment that turns the host synthetic cell that is susceptible to infection into cells that cannot fuse with a pathogen. The synthetic cell fusion system used throughout this paper relies on single-stranded DNA tags to induce liposome membrane fusion. To inoculate against fusion with a pathogen, we incubated the cells with DNA oligonucleotide complementary to the fusion oligo on the surface of the cells (Figure 6a). First, we needed to confirm that inoculation by incubation with complementary oligo works to abolish fusion. In the proof-of-principle experiments, two populations of liposomes were prepared: one population had TxTl with a plasmid encoding GFP under the control of the T7 promoter and the other population had TxTl and T7 RNA polymerase but no plasmid. After the fusion of these two populations, the GFP plasmid mixed with T7 RNA polymerase and induced GFP expression. If the two populations had single-stranded and complementary fusion tags, fusion occurred and GFP fluorescence was detected. If one or both populations were incubated with an oligonucleotide complementary to the fusion tag, the liposome fusion did not occur and GFP was not expressed (Figure 6b). Having established the inoculation system, we proceeded to introduce inoculation to protect host synthetic cells in three of the infection scenarios described earlier in this work. First, we used the parasite scenario described in Figure 2. The host cells, containing a GFP genome, were inoculated with an oligo complementary to the fusion tag on the surface of the host, and the host was mixed with the mCherry parasite. We mixed the host with the parasite at different ratios. Under all infection conditions, the parasite mCherry fluorescence was undetectable if the host was inoculated and the GFP fluorescence of the inoculated host did not decrease. The control experiments with the noninoculated host showed a decrease in the host GFP expression and an increase in the parasite mCherry signal, as expected (Figure 6c). Next, we applied the inoculation system to the case of a host infected with a pathogen carrying a genome-encoding MunI restriction enzyme capable of digesting the hosts’ GFP genome, like in experiments shown in Figure 3. The inoculated host did not show a decrease in GFP fluorescence after mixing with increasing amounts of the pathogen, while the noninoculated host was infected and the host GFP fluorescence decreased proportionally to the increasing amount of pathogen (Figure 6d). Inoculation was also effective against the aHL pathogen. The inoculated hosts showed no decrease in fluorescence upon mixing with the pathogen, while the noninoculated host’s GFP fluorescence decreased significantly with infection (Figure 6e). In all cases, this simple inoculation procedure, incubating the host with a DNA oligo complementary to the fusion oligo on the host’s surface, was sufficient to abolish the ability of the pathogenic synthetic cells to infect the host cells. Therefore, we demonstrated a simple physicochemical model of intervention that protects host cells from infection.

cistanche benefits for men-strengthen immune system
CONCLUSIONS
In this paper, we demonstrate methods for using synthetic minimal cells to model parasite and pathogenic infections. The motivation behind our work was twofold. We wanted to engineer complex, life-like behavior into synthetic cells, facilitating the goal of engineering synthetic cells that more closely resemble natural live cells. We also wanted to take advantage of the simple and controllable synthetic cell system to model the basic properties of infection, inoculation, and resistance processes. While synthetic cells do not evolve or self-replicate, this system allows for studying infection dynamics in a simple and easy-to-analyze model. We hope the principles of infection described here will help to create practical and applicable models for different pathogens and infection mitigation. Studying this simplified synthetic cell system highlighted the fundamental interactions between the parasite and the pathogen. While synthetic cells are not (yet) alive, and they do not have the ability to undergo Darwinian evolution, this system allows us to observe pathogen and host interactions without the ability to develop natural immunity or change the virulence of the pathogen. While the host infection dynamics presented here utilize bacteria-based synthetic cells, this system is not inherently limited to bacterial TxTl. It would even be possible to imagine using the system presented here to model infection dynamics in populations resembling natural ecosystems, taking advantage of the simplicity inherent to synthetic cells to fully control and study the interactions between different members of large populations.
REFERENCES
(1)Applications. Mater. Today 2016, 19, 516−532.
(2) Gaut, N. J.; Adamala, K. P. Reconstituting Natural Cell Elements in Synthetic Cells. Adv. Biol. 2021, 5, No. 2000188.
(3) Tan, C.; Saurabh, S.; Bruchez, M. P.; Schwartz, R.; Leduc, P. Molecular Crowding Shapes Gene Expression in Synthetic Cellular Nanosystems. Nat. Nanotechnol. 2013, 8, 602−608.
(4) Glantz, S. T.; Berlew, E. E.; Chow, B. Y. Synthetic Cell-like Membrane Interfaces for Probing Dynamic Protein-Lipid Interactions, 1st ed.; Elsevier Inc, 2019; Vol. 622.
(5) Zhao, J.; Zhang, Y.; Zhang, X.; Li, C.; Du, H.; Sønderskov, S. M.; Mu, W.; Dong, M.; Han, X. Mimicking Cellular Metabolism in Artificial Cells: Universal Molecule Transport across the Membrane through Vesicle Fusion. Anal. Chem. 2022, 94, 3811−3818.
(6) Damiano, L.; Stano, P. On the “Life-Likeness” of Synthetic Cells. Front. Bioeng. Biotechnol. 2020, 8, No. 953.
(7) Salehi-Reyhani, A.; Ces, O.; Elani, Y. Artificial Cell Mimics as Simplified Models for the Study of Cell Biology. Exp. Biol. Med. 2017, 242, 1309−1317.
(8) Chakraborty, T.; Wegner, S. V. Cell to Cell Signaling through Light in Artificial Cell Communities: Glowing Predator Lures Prey. ACS Nano 2021, 15, 9434−9444.
(9) Dupin, A.; Simmel, F. C. Signalling and Differentiation in Emulsion-Based Multi-Compartmentalized in Vitro Gene Circuits. Nat. Chem. 2019, 11, 32−39.
(10) Toparlak, O. D.; Zasso, J.; Bridi, S.; Serra, M. D.; Macchi, P.; Conti, L.; Baudet, M.-L.; Mansy, S. S. Artificial Cells Drive Neural Differentiation. Sci. Adv. 2020, 6, No. eabb4920.
(11) Robinson, A. O.; Venero, O. M.; Adamala, K. P. Toward Synthetic Life: Biomimetic Synthetic Cell Communication. Curr. Opin. Chem. Biol. 2021, 64, 165−173.
(12) Smith, J. M.; Chowdhry, R.; Booth, M. J. Controlling Synthetic Cell-Cell Communication. Front. Mol. Biosci. 2022, 8, No. 1321.
(13) Elani, Y. Interfacing Living and Synthetic Cells as an Emerging Frontier in Synthetic Biology. Angew. Chem., Int. Ed. 2021, 60, 5602− 5611.
(14) Rustad, M.; Eastlund, A.; Jardine, P.; Noireaux, V. Cell-Free TXTL Synthesis of Infectious Bacteriophage T4 in a Single Test Tube Reaction. Synth. Biol. 2018, 3, No. ysy002.
(15) Marshall, R.; Noireaux, V. Quantitative Modeling of Transcription and Translation of an All-E. coli Cell-Free System. Sci. Rep. 2019, 9, No. 11980.
(16) Garamella, J.; Marshall, R.; Rustad, M.; Noireaux, V. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. ACS Synth. Biol. 2016, 5, 344−355.
(17) Deich, C.; Cash, B.; Sato, W.; Sharon, J.; Aufdembrink, L.; Gaut, N. J.; Heili, J.; Stokes, K.; Engelhart, A. E.; Adamala, K. P. T7Max Transcription System. 2021, 1−28.
(18) Gaut, N. J.; Gomez-Garcia, J.; Heili, J. M.; Cash, B.; Han, Q.; Engelhart, A. E.; Adamala, K. P. Programmable Fusion and Differentiation of Synthetic Minimal Cells. ACS Synth. Biol. 2022, 11, 855− 866.
(19) Stengel, G.; Zahn, R.; Höök, F. DNA-Induced Programmable Fusion of Phospholipid Vesicles. J. Am. Chem. Soc. 2007, 129, 9584− 9585.
(20) Chizzolini, F.; Forlin, M.; Yeh Martín, N.; Berloffa, G.; Cecchi, D.; Mansy, S. S. Cell-Free Translation Is More Variable than Transcription. ACS Synth. Biol. 2017, 6, 638−647.
(21) Shin, J.; Noireaux, V. An E. coli Cell-Free Expression Toolbox: Application to Synthetic Gene Circuits and Artificial Cells. ACS Synth. Biol. 2012, 1, 29−41.
(22) Noireaux, V.; Libchaber, A. A Vesicle Bioreactor as a Step toward an Artificial Cell Assembly. Proc. Natl. Acad. Sci. U.S.A. 2004, 101, 17669−17674.
(23) Adamala, K. P.; Martin-Alarcon, D. A.; Guthrie-Honea, K. R.; Boyden, E. S. Engineering Genetic Circuit Interactions within and between Synthetic Minimal Cells. Nat. Chem. 2017, 9, 431−439.
(24) Noireaux, V.; Bar-Ziv, R.; Godefroy, J.; Salman, H.; Libchaber, A. Toward an Artificial Cell Based on Gene Expression in Vesicles. Phys. Biol. 2005, 2, P1−P8.
(25) Krinsky, N.; Kaduri, M.; Zinger, A.; Shainsky-Roitman, J.; Goldfeder, M.; Benhar, I.; Hershkovitz, D.; Schroeder, A. Synthetic Cells Synthesize Therapeutic Proteins inside Tumors. Adv. Healthcare Mater. 2018, 7, No. 1701163.
(26) Kwon, Y. C.; Jewett, M. C. High-Throughput Preparation Methods of Crude Extract for Robust Cell-Free Protein Synthesis. Sci. Rep. 2015, 5, No. 8663.
(27) Sun, Z. Z.; Hayes, C. A.; Shin, J.; Caschera, F.; Murray, R. M.; Noireaux, V. Protocols for Implementing an Escherichia Coli Based TXTL Cell-Free Expression System for Synthetic Biology. J. Vis. Exp. 2013, 79, 1−15.
(28) Fujii, S.; Matsuura, T.; Sunami, T.; Nishikawa, T.; Kazuta, Y.; Yomo, T. Liposome Display for in Vitro Selection and Evolution of Membrane Proteins. Nat. Protoc. 2014, 9, 1578−1591.
(29) Löffler, P. M. G.; Ries, O.; Rabe, A.; Okholm, A. H.; Thomsen, R. P.; Kjems, J.; Vogel, S. A DNA-Programmed Liposome Fusion Cascade. Angew. Chem., Int. Ed. 2017, 56, 13228−13231.






