Deep Structured Residual Encoder-decoder Network With A Novel Loss Function For Nuclei Segmentation Of Kidney And Breast Histopathology Images
Jul 11, 2023
Abstract
To improve the process of diagnosis and treatment of cancer disease, automatic segmentation of hematoxylin and eosin (H & E) stained cell nuclei from histopathology images is the first step in digital pathology. The proposed deep structured residual encoder-decoder network (DSREDN) focuses on two aspects: first, it effectively utilized residual connections throughout the network and provides a wide and deep encoder-decoder path, which results to capture relevant context and more localized features. Second, vanished boundary of detected nuclei is addressed by proposing an efficient loss function that better trains our proposed model and reduces the false prediction which is undesirable, especially in healthcare applications. The proposed architecture experimented on three different publicly available H&E stained histopathological datasets namely: (I) Kidney (RCC) (II) Triple Negative Breast Cancer (TNBC) (III) MoNuSeg-2018. We have considered F1-score, Aggregated Jaccard Index (AJI), the total number of parameters, and FLOPs (Floating point operations), which are mostly preferred performance measure metrics for comparison of nuclei segmentation. The evaluated score of nuclei segmentation indicated that the proposed architecture achieved a considerable margin over five state-of-the-art deep learning models on three different histopathology datasets. Visual segmentation results show that the proposed DSREDN model accurately segments the nuclear regions more than those of the state-of-the-art methods.
Keywords
Kidney cancer diagnosis and prognosis · Nuclei segmentation · Residual learning · Histopathology images.

Click here to know the effects of Cistanche
Introduction
Recent research trends show that the deep learning framework performed very well for segmentation, detection, and other computer vision tasks. In the last decade, with the advancement of new types of computation systems, proper strategies to handle overfitting problems to train very deep networks, and lots of changes that are suitable for deep learning networks. Segmentation of hematoxylin and eosin (H & E) from stained histopathology images is the primary prerequisite in artificial pathology. The histopathology slide preparations are discussed by Slaoui M et al. in [27], by the following steps: (I) Tissue collection (II) Fixation (III) Embedding (IV) Sectioning (V) De-paraffining (VI) Staining (VII) Digitizing the slide by whole slide imaging (WSI). There are several tissue collection methods which are fine needle aspiration, biopsy needle, excisional biopsy, etc. A larger biopsy has more information than a small needle biopsy because it preserves the large cellular context of histopathology slides. Fixation of tissue is needed for chemical and physical stabilization. Embedding is required to give a particular shape to the tissue such that it can be easily cut by the machines. Sectioning is required to get all three-dimensional tissue information in the form of many thin slides and two-dimensional information. Removing paraffin from the sectioned tissue is important, without de-paraffining the tissue may look a little bit blurry in some of the portions. Staining of the tissue slides is required because it is not visible or kind of transparent under bright field microscopy. The most widely used stains for histopathology images are hematoxylin and eosin. Segmentation tasks can be categorized into traditional or handcrafted feature extraction techniques and CNN-based deep learning approaches. Traditional segmentation methods are mostly based on similarity-based approach, discontinuity-based approach, watershed techniques, active contour methods and their variants, superpixel, and clustering-based methods, etc. The similarity-based approach discussed by Gonzalez R C et al. in [8], is based on local thresholding, global thresholding, adaptive thresholding, Otsu’s thresholding, region growing, region splitting, and merging, where these methods try to group and segment similar pixels. For image histograms having flat valleys, the similarity-based approach does not work well and the wrong selection of threshold value may result in over-segmentation and under-segmentation in this case. The discontinuity-based approach tries to segment those pixels which are isolated in some manner like points, lines, and edges, and it is a mask processing-based approach. This method requires different operators at different stages. Cousty J et al. proposed a watershed segmentation method in [4], based on the split, merge, and marker-controlled watershed. Detected boundaries in the watershed method depend on cell complexity. Song T et al. proposed active contour segmentation in [28], where they consider intensity information and local edge information for the detection of object boundaries. The superpixel segmentation method used by Albayrak A et al. in [1], is based on the cluster of connected pixels having identical features. It considers the color and coordinates information of neighbor pixels. This technique provides better regional information but is not very effective in the case of cell segmentation. Clustering-based segmentation proposed by Win K Y et al. in [37], performs grouping based on their similarity. In recent research work, most of the authors reported that the segmentation technique based on a deep convolutional neural network performs far better than the conventional segmentation approach. A concise review of CNN based approach is presented in Section 2. Deep learning segmentation methods also suffer from many challenges. If we categorize these challenges, it will come under the following aspects.
1. Due to large variations of tissue appearance and a varied spectrum of class and sub-class of tissues, it is difficult to recognize.
2. Segmentation of complex boundaries, overlapped boundaries, and vanishing boundaries is not an easy task.
3. Preparation of ground truth in the case of supervised learning is also a big challenge. Supervision of experienced pathologists is necessary since prediction accuracy depends on annotated ground truth.
In the case of complex histopathology images, conventional methods suffer either over-segmentation or under-segmentation. The proposed approach focuses to separate the overlapped and vanished nuclear regions from histopathology images. To address the challenges in the segmentation of nuclei from histopathology images our contributions in this paper are as follows.
1. To strengthen the multi-level intermediate features, our proposed DSREDN model effectively utilized the strength of residual learning.
2. Through empirical evidence and careful experimentation and analysis, we proposed a novel loss function. Visual results and performance matrices indicate that our loss function better trains the model and accurately segments the nuclear regions compared to the state-of-the-art methods.

Cistanche tubulosa
Related work
Most of the CNN architecture for the cell segmentation task consists of an encoder-decoder path for feature extraction. Much of the recent research utilizes lots of potential opportunities like improving training strategies, handling overfitting problems, better optimization methods, and many other strategies to obtain better prediction accuracy. However, many authors reported their result which is very efficient but an accurate and efficient segmentation algorithm is still open-ended research due to the complexity of histopathology images. One of the significant contributions by Ronneberger et al. in [26], called UNet, provides a very good direction and a dramatic breakthrough in the field of biomedical image segmentation. UNet is a symmetric encoder-decoder convolutional network and has a large number of feature channels that allow to extract features to the higher layer in a deep network. Repeated application of (3 x 3) convolution kernel followed by ReLU activation, (2 x 2) max-pooling and (2 x 2) up-sampling with stride size of 2, and (1 x 1) convolution followed by sigmoid activation at final layer, a total of 23 layers in the network. In [36], Veit A et al. realized through their experiment that if a network has a collection of paths then a shorter path is enough during training, or a very deep path is not required during training. These multiple paths do not strongly depend on each other and their smooth co-relation with multiple valid paths increases the performance of the network. In [22], Milletari F et al. proposed an encoder-decoder convolutional network for three-dimensional data by utilizing dice loss as a loss function. Their empirical evaluation achieves better performance on the strong imbalance dataset. In [24], Nogues I et al. proposed an architecture for the detection of lymph nodes by two fully nested supervised convolutional networks and a structured conditional random field optimization strategy. Degradation of information in a deeper network was addressed by Kaiming He et al. in [9], by introducing a deep residual network which is easier to train and optimize. The residual connection is realized by skipping one or more layers to restore the flow of information in a deep network. For segmentation and detection of histological objects, Chen H et al. in [5], introduced a contour-aware model that extracts multi-level information under auxiliary supervision. In [10], Huang G et al. proposed a convolutional network, which strengthens the overall flow of the input feature map by feeding the preceding layer input as well as the original input. Their experiment also indicates that due to the integration of identity mapping, the model learns more compact features and reduces the vanishing gradient problem. In the case of an imbalanced dataset, predictions are biased towards high precision and low recall which is not tolerable, especially in the medical field. This problem is addressed by Salehi S M et al. in [29], which trained the deep network, even with a highly imbalanced dataset, and handled effectively where false negative prediction is much more dangerous than false positive. The behavior of loss functions such as weighted cross-entropy and dice loss with different learning rates examined by Sudre C H et al. in [30], on medical images and house datasets. Their experiment found that as the level of imbalance increases overlap measure-based loss function is more effective. A very efficient in terms of memory and time for semantic segmentation of road and indoor scenes, an encoder-decoder architecture called SegNet by Badrinarayanan V et al. in [3]. SegNet generates a sparse feature decoder that upsamples with the transferred pool and its lower resolution input from its encoder. To accurately segment near boundary regions Zhou S et al. [38], used a residual network with a dilated convolution block. They utilize many hierarchical blocks in parallel to retrieve meaningful semantic information. To handle class imbalance problems or reduce false-negative predictions in healthcare, Hashemi S R in [11] proposes a 3D-dense CNN with Tversky index-based asymmetric similarity loss that trains the network with the lowest surface distance. Complex boundary-related segmentation problem addressed by Naylor P et al. in [25], by formulating a loss function based on intra- nuclear distance. Their encoder-decoder model outperform FCN, FCN+PP, Mask R-CNN, U-Net, and U-Net+PP experimented with TNBC and MoNuSeg datasets. Meaningful extensions in standard encoder-decoder by incorporating an additional module called attention gate by Schlemper J et al. in [31], and attention as well as the residual mechanism by Lal S et al. in [20], where the network is trained in such a way that it suppresses irrelevant features while highlighting the meaningful feature. For road scene segmentation Malekijoo A et al. in [23], utilized the autoencoder-based model where convolution, deconvolution, and pyramid pooling were applied for reinforcing the local feature. For the segmentation of microscopic, MR, and CT images an encoder-decoder architecture by Zhou S et al. in [39], linked meaningful connections to precisely locate the complex boundaries. For the segmentation of nuclei in pathology images, the Lal S et al. model [21], consists of adaptive color deconvolution, multiscale thresholding followed by morphological operations, and other post-processing steps. For the segmentation of medical images, a novel loss function by Karimi D et al. in [16], estimated Hausdorff distance using the morphological operation method, distance transformation method, and circularly convoluted kernels of different radii. Utilizing methods of reduction of the Hausdorff distance, they train CNN for various microscopy images and compare their results with a commonly used loss function. Hanif M S et al. in [12], proposed a competitive residual network by stacking multiple residual units called a wide network. Their study concluded that the performance of such a wide network is better than the deep and thin network. Chanchal A K et al. and Aatresh A A et al. in [2, 6], used separable convolution pyramid pooling and dimension-wise pyramid pooling for nuclei segmentation tasks.

Cistanche capsules
Proposed architecture
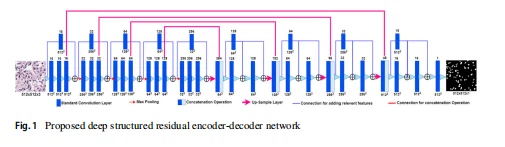
For the segmentation of microscopy images, an encoder-decoder architecture is best suited because if an encoder has regular convolution layers and max-pooling layers, then it captures the context in the image very effectively. The decoder path presents the output by gradually applying up-sampling, collecting relevant features from the encoder, and enabling precise localization. Each of the filters in the encoder side of the DSREDN network shown in Fig. 1, accepts input of flexible size. we have applied regular (3 x 3) 2D standard convolution, batch normalization, and max-pooling. To avoid saturation problems and loss of information while going deeper into the network, we restored the lower-layered information by creating an additional path parallel to the main path of the network. These two paths are not strongly correlated to each other and it avoids vanishing gradient problems. For each of the filter sizes, the entire encoder side of the DSREDN network consists of three convolution layers in parallel with a single convoluted path that focused to flow the more contextual feature in the network. Since the effectiveness of the decoder path to generate the final output depends on the collection of contextual features from the encoder side, we have a slightly different path on the decoder side, for the optimal processing of the collected feature. By this procedure, our DSREDN network becomes wide and deep instead of thin and deep. DSREDN network trained with RGB images of size (512 x 512 x 3). Five stages of the encoder path having five different filter sizes and corresponding decoder path consist of (a) 2D convolution of kernel size (3 x 3) with ReLU activation (b) A high-resolution layer (c) (2 x 2) max-pooling layer in encoder path to reduce the spatial size of the image and corresponding (2 x 2) up-sampling layer in decoder side to collect contextual feature from encoder side by concatenation operation (d) At the final stage a (1 x 1) convolution is used to map the size (512 x 512 x 16) to (512 x 512 x 1) with sigmoid activation.
Conclusion
This paper proposed a CNN-based architecture called a deep structured residual encoder-decoder network (DSREDN), that addressed two major concerns in automatic nuclei segmentation. The first major concern was to identify nuclei from histopathology images having a widely varied spectrum with a large number of artifacts. This problem was addressed by introducing a powerful encoder-decoder having two paths that have more discriminative capability and were able to retrieve relevant and compact textural information. The implemented networks effectively leverage the strength of residual learning as well as encoder-decoder architecture by incorporating wide and deep network paths that strengthen the intermediate features. We proposed an efficient loss function through careful experimentation and analysis to segment the nuclei having complex or vanishing boundaries which were the second major issue in the segmentation task. We have used the most preferred performance matrices F1-score and AJI score by performing experiments on the three different publicly available H&E stained histopathological datasets. The obtained quality metrics and predicted nuclear regions of the proposed framework were better in comparison to those of the state-of-the-art models.

Cistanche pills
Although the proposed model produced excellent results, the feature space may be enriched further by incorporating a high-performance feature extraction module. Also, the proposed method can be generalized to work on more image modalities. This study is a binary segmentation of histopathology images, here we can only segment the nuclear regions. In the future, we can grade these nuclear regions into their sub-types. Few innovative applications of different image modalities were reported by Shoeibi A et al. in [32, 33], in which generative adversarial networks (GANs), recurrent neural networks (RNNs), autoencoders (AEs), convolutional neural networks (CNNs), deep neural networks (DNNs), and other hybrid networks have been developed for automated detection of COVID-19 and multiple sclerosis. In [18, 34], Khodatars M et al. and Sadeghi D et al. illustrated the applicability of deep learning for the diagnosis of autism spectrum disorder and schizophrenia disease detection. These examples highlight how the field of computer-aided diagnosis systems is changing rapidly, and that there may still be numerous applications that have not been focused on yet.
How Cistanchis Enhance Kidney Function
Cistanche is a medicinal herb that has long been used in traditional Chinese medicine to enhance kidney function. It is believed to have various benefits due to its active components, such as phenylethanoid glycosides and iridoids.
Studies suggest that Cistanche may promote kidney function by improving renal blood flow, reducing oxidative stress, and enhancing the production of growth factors that support kidney health. Additionally, it may help regulate blood pressure and reduce inflammation, which are important factors for kidney health.
Moreover, Cistanche has shown the potential in protecting against kidney damage caused by certain medications or toxins. It may have a protective effect on the kidneys by inhibiting inflammatory responses and reducing cell death.
However, it's important to note that further research is needed to fully understand the mechanisms and effectiveness of Cistanche in enhancing kidney function. As with any herbal remedy, it's advisable to consult with a qualified healthcare professional before using it for medicinal purposes.
References
1. Albayrak A, Bilgin G (2019) Automatic cell segmentation in histopathological images via two-staged superpixel-based algorithms. Med Biol Eng Comput 57(3):653–665
2. Aatresh AA, Yatgiri RP, Chanchal AK, Kumar A, Ravi A, Das D, Raghavendra BS, Lal S, Kini J (2021) Efficient deep learning architecture with dimension-wise pyramid pooling for nuclei segmentation of histopathology images. Comput Med Imaging Graph 93:101975. https://doi.org/10.1016/j.compmedimag.2021.101975
3. Badrinarayanan V, Kendall A, Cipolla R (2017) Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans Pattern Anal Mach Intell 39(12):2481–2495
4. Cousty J, Bertrand G, Najman L, Couprie M (2010) Watershed cuts: Thinnings, shortest path forests, and topological watersheds. IEEE Trans Pattern Anal Mach Intell 32(5):925–939
5. Chen H, Qi X, Yu L, Heng PA (2016) DCAN: deep contour-aware networks for accurate gland segmentation. Computer Vision and Pattern Recognition. arXiv:1604.02677v1 [cs.CV]
6. Chanchal AK, Kumar A, Lal S, Kini J (2021) Efficient and robust deep learning architecture for segmentation of kidney and breast histopathology images. Comput Electr Eng 92:107177. https://doi.org/10.1016/j.compeleceng.2021.107177
7. Chanchal AK, Lal S, Kini J (2021) High-resolution deep transferred ASPPU-Net for nuclei segmentation of histopathology images. Int J Comput Assist Radiol Surg. https://doi.org/10.1007/s11548-021-02497-9
8. Gonzalez RC, Woods RE (2006) Digital image processing, 3rd ed. Prentice Hall, New York, USA. ISBN-013168728X
9. He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. 2016 IEEE Conference on computer vision and pattern recognition (CVPR), Las Vegas, NV. pp 770–778. https://doi.org/10.1109/CVPR.2016.90
10. Huang G, Liu Z, Maaten L, Weinberger KQ (2017) Densely connected convolutional networks. IEEE Conference on computer vision and pattern recognition (CVPR), Honolulu. pp 2261–2269. https://doi.org/10.1109/CVPR.2017.243
11. Hashemi SR, Salehi SM, Erdogmus D, Prabhu SP, Warfield SK, Gholipour A (2019) Asymmetric loss functions and deep densely-connected networks for highly-imbalanced medical image segmentation: application to multiple sclerosis lesion detection. In: IEEE Access, vol 7, pp 1721–1735. https://doi.org/10.1109/ACCESS.2018.2886371
12. Hanif MS, Bilal M (2020) Competitive residual neural network for image classification. ICT Express 6(1):28–37. https://doi.org/10.1016/j.icte.2019.06.001
13. Ioffe S, Szegedy C (2015) Batch normalization: accelerating deep network training by reducing internal covariate shift. Machine Learning. arXiv:1502.03167
14. Irshad H, Kouhsari LM, Waltz G, Bucur O, Nowak JA, Dong F, Knoblauch NW, Beck AH (2015) Crowdsourcing image annotation for nucleus detection and segmentation in computational pathology: evaluating experts, automated methods, and the crowd. In: Pacific Symposium on biocomputing (PSB), pp 294–305. https://doi.org/10.13140/2.1.4067.0721
15. Jadon S (2020) A survey of loss functions for semantic segmentation. [Online]. Available: arXiv:2006.14822
16. Karimi D, Salcudean SE (2020) Reducing the Hausdorff distance in medical image segmentation with convolutional neural networks. IEEE Trans Med Imaging 39(2):499–513
17. Kumar N, Verma R, Sharma S, Bhargava S, Vahadane A, Sethi A (2017) A dataset and a technique for generalized nuclear segmentation for computational pathology. IEEE Trans Med Imaging 36(7):1550– 1560
18. Khodatars M, Shoeibi A, Sadeghi D, Ghaasem N, Jafari M, Meridian P, Khadem A, Alizadehsani R, Zare A, Kong Y, Khosravi A, Nahavandi S, Hussain S, Acharya UR, Berk M (2021) Deep learning for neuroimaging-based diagnosis and rehabilitation of Autism Spectrum Disorder: A review. Comput Biol Med 139:104949. https://doi.org/10.1016/j.compbiomed.2021.104949
19. Kingma DP, Ba J (2015) Adam: A method for stochastic optimization. In: International conference on learning representations, vol 9. arXiv:1412.6980v9 [cs.LG]
20. Lal S, Das D, Alabhya K, Kanfade A, Kumar A, Kini J (2021) NucleiSegNet: Robust deep learning architecture for the nuclei segmentation of liver cancer histopathology images. Comput Biol Med 128:104075
21. Lal S, Kanfade A, Alabhya K, Dsouza R, Kumar A, Chanchal AK, Maneesh M, Peryail G, Kini J (2020) A robust method for nuclei segmentation of H&E stained histopathology images. 7th IEEE international conference on signal processing and integrated networks (SPIN2020), Amity University Delhi NCR, Noida, UP
22. Milletari F, Navab N, Ahmadi SA (2016) V-Net: fully convolutional neural networks for volumetric medical image segmentation, fourth international conference on 3D vision (3DV). Stanford, CA. pp 565–571. https://doi.org/10.1109/3DV.2016.79
23. Malekijoo A, Fadaeieslam MJ (2019) Convolution-deconvolution architecture with the pyramid pooling module for semantic segmentation. Multimed Tools Appl 78:32379–32392. https://doi.org/10.1007/s11042-019-07990-7
24. Nogues I et al (2016) Automatic lymph node cluster segmentation using holistically-nested neural networks and structured optimization in CT images. In: Medical image computing and computer assisted intervention – MICCAI 2016. Lecture notes in computer science, vol 9901. Springer, Cham. https://doi.org/10.1007/978-3-319-46723-845
25. Naylor P, Lae M, Reyal F, Walter T (2019) Segmentation of nuclei in histopathology images by deep regression of the distance map. IEEE Trans Med Imaging 38(2):448–459
26. Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional networks for biomedical image segmentation. In: Proc. MICCAI. Springer, Munich, Germany, pp 234–241
27. Slaoui M, Fiette L (2011) Histopathology procedures: from tissue sampling to histopathological evaluation. Methods Mol Biol (Methods Protoc) 691:69–82
28. Song T, Sanchez V, EIDaly H, Rajpoot NM (2017) Dual-channel active contour model for megakaryocytic cell segmentation in bone marrow trephine histology images. IEEE Trans Biomed Eng 64(12):2913–2923
29. Salehi SM, Erdogmus D, Gholipour A (2017) Tversky loss function for image segmentation using 3D fully convolutional deep networks. In: Proc, Int Workshop Mach Learn Med Imag. Springer, Cham, Switzerland, pp 379–387
30. Sudre CH, Li W, Vercauteren T, Ourselin S, Cardoso MJ (2017) Generalised dice overlap as a deep learning loss function for highly unbalanced segmentations. In: Deep learning in medical image analysis and multimodal learning for clinical decision support. Springer, pp 240–248
31. Schlemper J, Oktay O, Schaap M, Heinrich M, Kainz B, Glocker B, Rueckert D (2019) Attention gated networks: Learning to leverage salient regions in medical images. Med Image Anal 53(ISSN 1361- 8415):197–207
32. Shoeibi A, Khodatars M, Alizadehsani R, Ghassemi N, Jafari M, Meridian P, Khadem A, Sadeghi D, Hussain S, Zare A, Sani ZA, Bazeli J, Khozeimeh F, Khosravi A, Nahavandi S, Acharya UR, Shi P (2020) Automated detection and forecasting of covid-19 using deep learning techniques: a review. Machine Learning. arXiv:2007.10785 [cs.LG]
33. Shoeibi A, Khodatars M, Jafari M, Meridian P, Rezaei M, Alizadehsani R, Khozeimeh F, Gorriz JM, Heras J, Panahiazar M (2021) Applications of deep learning techniques for automated multiple sclerosis detection using magnetic resonance imaging: A review, Image, and Video Processing. arXiv:2105.04881
34. Sadeghi D, Shoeibi A, Ghassemi N, Meridian P, Khadem A, Alizadehsani R, Teshnehlab M, Gorriz JM, Nahavandi S (2021) An overview on artificial intelligence techniques for the diagnosis of schizophrenia based on magnetic resonance imaging modalities: methods, challenges, and future works. Machine Learning. arXiv:2103.03081
35. Sugino T, Kawase T et al (2021) Loss weightings for improving imbalanced brain structure segmentation using fully convolutional networks, healthcare. MDPI 9(8):938
36. Veit A, Wilber M, Belongie S (2016) Residual networks behave like ensembles of relatively shallow networks. Neural Inf Process Syst pp 550–558. arXiv:1605.06431
37. Win KY, Choomchuay S, Hamamoto K (2017) K mean clustering based automated segmentation of overlapping cell nuclei in pleural effusion cytology images. International conference on advanced technologies for communications (ATC). pp 265–269. https://doi.org/10.1109/ATC.2017.8167630
38. Zhou S, Nie D, Adeli E, Gao Y, Wang L, Yin J, Shen D (2018) Fine-grained segmentation using hierarchical dilated neural networks. In: Medical image computing and computer assisted intervention, vol 11073. Springer, Cham, pp 488-496
39. Zhou S, Nie D, Adeli E, Yin J, Lian J, Shen D (2020). In: IEEE Transactions on Image Processing, vol 29, pp 461–475. https://doi.org/10.1109/TIP.2019.2919937
Amit Kumar Chanchal 1 · Shyam Lal 1 ·Jyoti Kini 2
1 Department of Electronics and Communication Engineering, National Institute of Technology Karnataka, Surathkal, Mangaluru-575025, Karnataka, India
2 Department of Pathology, Kasturba Medical College Mangalore, Manipal Academy of Higher Education, Manipal, India






