Part Ⅱ Metabolic Understanding Of The Genetic Dysregulation in The Tumor Microenvironment Of Kidney Renal Clear Cell Carcinoma
May 08, 2023
Result
1. Identification of Differentially Expressed Metabolic Genes in KIRC.
To explore the metabolic dysregulation in KIRC, we explored the available TCGA data to get deep insights into metabolism-targeted therapeutics in the clinic. For this purpose, we selected a set of 1916 metabolic genes that were intersected from two different datasets [16, 17] and screened out 1100 differentially expressed genes in tumor vs normal tissues (Supplementary Table 2). These differentially expressed metabolic genes were plotted in volcanoes and heat maps (Figures 1(a) and 1(b)). Out of 1100 differentially expressed metabolic genes, 78 genes were upregulated and 163 genes were downregulated. Moreover, there were 859 genes were unchanged. The heat map represents the individual expression index of those differential metabolic genes in tumor and normal samples (Figure 1(b)). Next, we identified the top 10 differentially expressed metabolic genes; among them, ENPP3, NNMT, CYP2J2, SCD, and HK2 were upregulated and HSD11B2, HMGCS2, HPD, HS6ST2, and ALDOB were downregulated. The box plots of these DEmGs are shown in Figure 1(c). Among the upregulated genes, ENPP3 is ~7-fold expressed in tumors. Alternatively, the gene ALDOB is ~5-fold downregulated in analyzed tumor samples.
In addition, we evaluated the KEGG pathway and GO analyses of DEmGs. KEGG pathway analysis revealed that the upregulated genes were significantly enriched in carbon metabolism, HIF1 signaling, and glycolysis/gluconeogenesis with a higher gene ratio (8–9 number of genes in each pathway) (Figure 1(d)). Similarly, among downregulated DEmGs, we found that carbon metabolism and valine, leucine, and isoleucine degradation were the top pathways affected by metabolically active genes (Figure 1(e)). The pathways related to peroxisome organelle were also significantly enriched in a downregulated group of gene tumor samples. Noteworthy, compared with those in the upregulated genes, the pathways involved in downregulated genes have higher significant p values. It is worth mentioning that most of the KEGG pathways enriched in downregulated gene categories were related to amino acid metabolism. To further dissect the involvement of DEmG in tumorigenesis, the GO functional analyses of upregulated and downregulated genes were performed. We divided the GO ontology into three functional subontology groups, BP (biological process), CC (cellular component), and MF (molecular function) (Figures 1(f) and 1(g)). In addition, GSEA analysis revealed a significant increase in the enrichment of genes associated with BENPORATH_MYC_TARGETS_WITH_EBOX in tumors, while BROWN_ MYELOID_CELL_DEVELOPMENT_UP, KEGG_ALPHA_ LINOLENIC_ACID_METABOLISM, and KEGG_ETHER_ LIPID_METABOLISM were found to be negatively enriched. In the next stage, we constructed a protein-protein interaction PPI network with up and downregulated DEmGs. A number of the genes showed interaction with each other. Through these genes’ interactions, we isolated hub genes. Each node is discrete from the other based on the degree value; further, we isolated the top 7 hub genes for PPI. We also explored the correlation between this hub gene expression and clinicopathological features of KIRC in TCGA datasets.

Click here to know what are the Cistanche benefits.
2. Network Analysis Reveals Basic Metabolic Changes in Various Tumor Ontologies.
Next, the expression data of DEmGs were selected and used as the input data for WGCNA, which identified 6 distinct coexpression modules containing a different number of genes for each module (Figure 2(a)). We correlated differential genes with external traits and identified the modules that were significantly associated with clinical traits (Figure 2(b)). Based on the correlation coefficient, we found that MEturquoise modules were negatively correlated with the survival status. GO and KEGG pathway enrichment analyses were performed using genes from these modules (Figures 2(c) and 2(d)). The most enriched KEGG pathways were valine, leucine, and isoleucine degradation; carbon metabolism; propanoate metabolism; fatty acid metabolism; fatty acid degradation; peroxisome and butanoate metabolism; glyoxylate and dicarboxylate metabolism; and tryptophan metabolism (Figure 2(c)). The genes related to BP terms were predominantly enriched in small molecule, carboxylic acid, and organic acid catabolic processes. The genes related to CC terms were mainly enriched in the mitochondrial matrix. The differentially expressed genes related to MF were mainly enriched in coenzyme binding (Figure 2(d)). In addition, we performed a survival analysis of 8 genes in the survival module. Patients with higher ACADSB, PANK1, SLC25A4, PCCA, HADH, AUH, ACAT1, and ALDH6A1 expression had a longer survival rate than those with lower expression of these genes (p = 0) (Figures 2(e)–2(l)).
3. Clustering of the KIRC Patients.
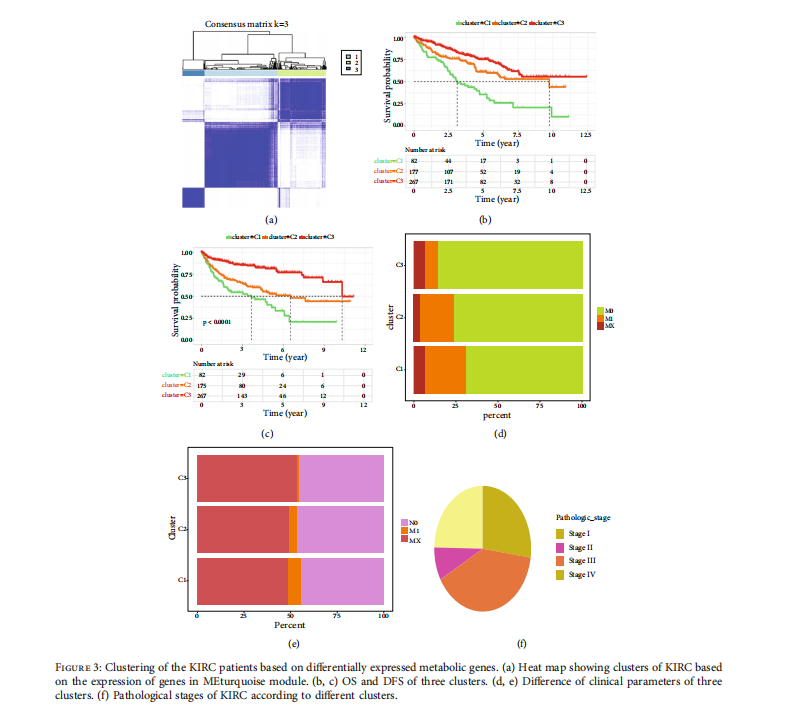
We selected top DEmGs for cluster analysis; the KIRC patients were grouped into three clusters based on the differential expression of metabolic genes. Figure 3(a) shows the heat maps of DEmGs in the KIRC patients. The color scale indicates the expression value (light blue indicates a lower expression value; darker blue indicates higher gene expression values).
KM curves were plotted to compare the overall survival of the three clusters for KIRC patients. The overall survival rates differed significantly across the three clusters (p < 0:01 Figure 3(b)). Cluster 1 showed a worse survival rate compared with Cluster 2 and Cluster 3. The PFS survival rate also differed significantly among the 3 clusters (p < 0:001, Figure 3(c)), and cluster 1 exhibited a worse PFS survival rate compared with cluster 2 and cluster 3.
Different colors in our model represent clinical parameters and underlying pathological stages (Figure 3(d)). Cluster 3 has lower Mo ratios and higher M1 value as compared with clusters 1 and 2 suggesting higher cancer metastasis and more advanced stage of tumors in cluster 3 than clusters 1 and 2. Similarly, in cluster 3, cancer has spread more to the lymph nodes (higher N1) as compared with those in clusters 1 and 2. Most of the KIRC patients were diagnosed at stages III and IV (Figures 3(e) and 3(f)), suggesting larger or expanded tumors, as well as moving through the blood or lymphatic system to a distant region in the body.
4. Immune Status of Three Clusters.
We used the ESTIMATE algorithm to estimate the stromal and immune scores of a series of KIRC tissues based on their metabolic transcriptional profiles (Figure 4(a)). Later, these scores were taken into account to develop a stromal-immune score-based metabolic genes signature for prognosis stratification in KIRC. As shown in Figure 4(a), three cluster groups (C1, C2, and C3) were stratified in box plots based on their stromal-immune score. Among the three clusters, C1 showed a higher significant score in both stromal and immune classifications.
Furthermore, three clusters were analyzed by CIBERSORT with a p-value < 0.1 (Figure 4(b)). The tumor purity, immune score, and stromal score along with the pathological stages of 3 clusters are shown at the top of the heat map. In this analysis, we majorly found that regulatory T cells (Tregs) were enriched in cluster C1 and patients in C1 were mainly at pathological stages III and IV. Moreover, the activated NK cells, CD8+ T cells, T follicular helper cells, and M0 macrophages in the C1 cluster; CD8+ T cells and T follicular helper cells in the C2 cluster; and resting mast cells, M2 macrophages, resting memory CD4 T cells, monocytes, naive B cells, and M1 macrophages in the C3 cluster were also detected (Figure 4(b)).
Apart from CIBERSORT, we employed other algorithmic packages to check the status of immune infiltration. The hierarchical heat map of MCP analysis is shown in Figure 4(c). The key findings of MCP analysis were neutrophil infiltration and endothelial cell infiltration in cluster C3 that was missing in cluster C1. This analysis also revealed NK cell, monocytic lineage, and myeloid dendritic cell infiltration in cluster C3. Other immune cell populations were mixed in three analyzed clusters (Figure 4(c)).
To complement CIBERSORT and MCP analyses, we applied ssGSEA to quantify infiltration levels for immune cell types implemented in R package GSVA. Three clusters’ data were fed to the ssGSEA package and obtained richness of 28 immune-related cells and types in KIRC samples. Results revealed that C1 and C2 had more immune infiltration; some innate immune cells, including NK, neutrophils, and eosinophils, were mixed in 3 clusters (Figure 4(d)).

5. Construction and Validation of the Predicting Model Based on DEmGs.
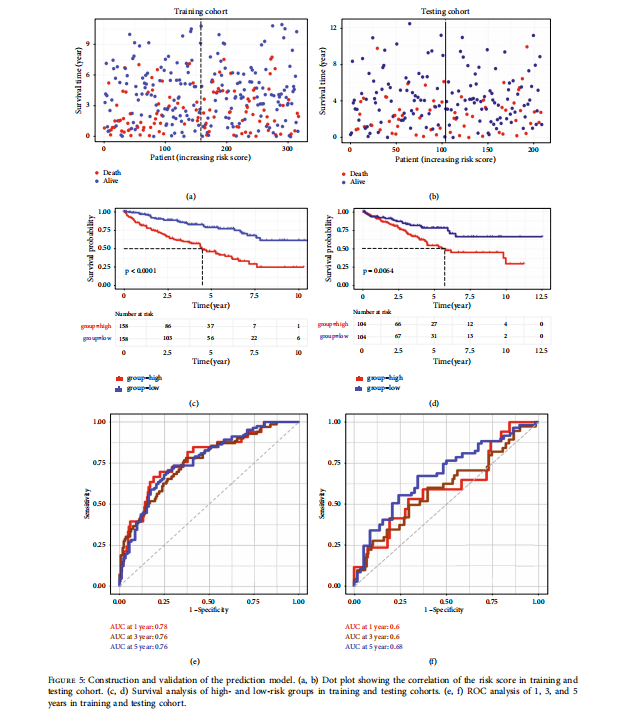
Lastly, we constructed and validated the prediction model based on the differential expression of the metabolic genes. We calculated the immune-related risk score of DEmGs based on overall survival. For this purpose, we devised two groups for evaluating the correlation of the risk score; one is for the training cohort, and the other for the testing cohort. We found that overall survival was low and scattered across the risk score (Figures 5(a) and 5(b)). Next, based on the median risk score, we assigned KIRC patients to high- and low-risk groups for further evaluation. We then performed a survival analysis of these two risk groups in training and testing cohorts. As expected, the high-risk groups were found to have a low survival as compared with the low-risk groups (Figures 5(c) and 5(d)). Furthermore, ROC curve analysis was performed for training and testing cohorts. We observed a ROC score of 0.68 at 5 years in testing cohorts, which indicates good performance in predicting the prognosis of KIRC (Figures 5(e) and 5(f)). In addition to the ROC curve analysis, we also executed the LASSO COX regression model to validate our prognostic model as indicated by partial likelihood deviance (Supplementary Figure 5(a)) and regression coefficient of the DEmGs (Supplementary Figure 5(b)). Lastly, five genes (ABCG1, CRYL1, FDX1, PANK1, and SLC44A) were predicted to be potential prognostic factors with HR < 1 (Supplementary Figure 5(c)).
6. Underlying Mechanisms of the KIRC Progression.
To investigate further the underlying mechanism for KIRC progression, we conducted differential expression analysis among all clusters and utilized a heat map plot to visualize the results (Figure 6(a)). To identify the signaling pathways of the DEmGs, we performed KEGG and GO enrichment analyses of the DEGs in three clusters. In short, these results revealed that DEGs of three clusters were mainly enriched in focal adhesion, the Foxo signaling pathway, and the Apelin signaling pathway for cluster C3 and mineral absorption, neutrophil extracellular trap formation, and staphylococcus aureus infection for cluster C2 (Figure 6(b)). In addition, GO functional analysis of DEGs uncovered MF-, CC-, and BP-related ontologies shown in Figure 6(c). Interestingly, the differential expression analysis disclosed abnormal behavior of genes’ regulation in three clusters. Mostly, NUDT1 was highly expressed in C1, which had the worst survival. Further investigation revealed that NUDT1 expression was significantly downregulated through the progression from C1 to C3 (Figure 6(d)). Moreover, the NUDT1 was found to be upregulated in KIRC tumor samples (Figure 6(e)). Next, we highlighted the NUDT1 expression in each tumor’s stages (Figure 6(f)). Overall survival analysis was also performed by the Kaplan-Meier plotter, and we found that patients with higher expression of NUDT1 had worse overall survival (HR = 1:82 (1.34–2.48), log-rank p = 0:00012) (Figure 6(g)).

Lastly, we performed KEGG and GO functional enrichment analyses for genes interacting with NUDT1. The genes were divided into two groups—positively correlated with NUDT1 and negatively correlated with NUDT1. KEGG pathway analysis showed that positively correlated genes were mainly enriched in the ribosomal pathway, Huntington's disease, amyotrophic lateral sclerosis, and Alzheimer’s disease. On the other hand, the negatively correlated genes were enriched mainly in hepatitis B and Foxo signaling (Figure 6(h)). Moreover, the GO ontology of three different groups MF, CC, and BP for both positively and negatively correlated genes was shown in Figure 6(i). In addition, we found that the expression of NUDT1 was highly correlated with the infiltration of immune cells (Supplementary Figure 6) and different clinical features of the KIRC patients (Table 1).

Cistanche supplements
7. Loss of NUDT1 Inhibits Renal Cancer Cell Proliferation and Migration.
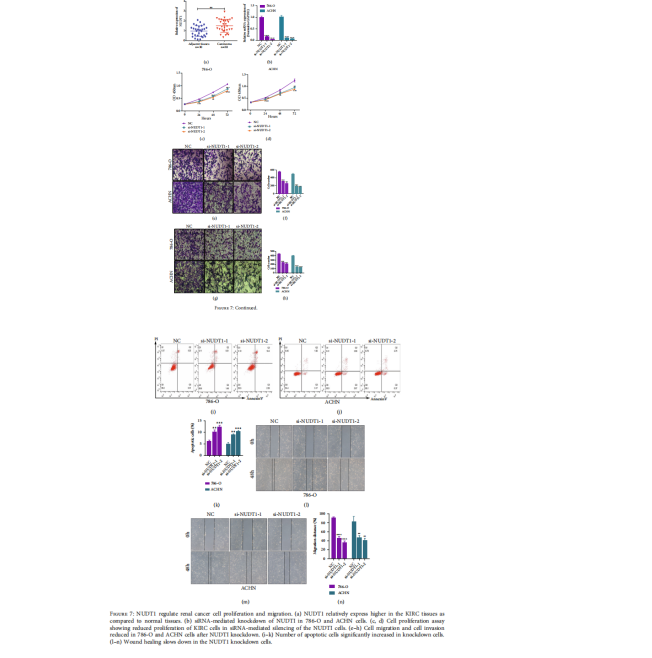
Next, we compared the expression level of NUDTI in KIRC tissues and their associated normal tissues, which revealed that NUDT1 is highly expressed in KIRC tissues (Figure 7(a)). Furthermore, we determined the effects of NUDT1 loss on renal cancer cell lines by using siRNA-mediated inhibition. NUDT1 was targeted for siRNA knockdown in two cell lines 786-O and ACHN and the NUDT1 mRNA levels were successfully inhibited as evidenced by qPCR analysis (Figure 7(b)). Following the siRNA-mediated knockdown of NUDT1, the cell viability assay showed reduced cell viability in both cell lines (Figures 7(c) and 7(d)). Afterward, cell migration assay upon knockdown of NUDT1 showed significantly reduced cell migration in NUDT1-depleted 786-O and ACHN cells (Figures 7(e) and 7(f)). The migration capacity of 786-O cells was reduced to about 50%, and a 70% reduction was observed in ACHN cells upon NUDT1 knockdown (Figure 7(f)). The cell invasion was also inhibited in both cell lines when the NUDT1 gene was knocked down (Figures 7(g) and 7(h)). To complement the migration, we also performed a wound healing assay when NUDT1 was depleted from 786-O and ACHN cell lines and we observed similar results of reduced wound healing capability in both cell lines lacking NUDT1 (Figures 7(l)–7(n)). Based on these results, we hypothesized that loss of NUDT1 could lead to apoptosis in renal cancer cells. Therefore, we measured the percentage of the apoptotic cells upon silencing of NUDT1. Interestingly, we discovered that the percentage of apoptotic cells was significantly increased in NUDT1-depleted cells (Figures 7(i)–7(k)).

Discussion
Kidney renal clear cell carcinoma (KIRC) is one of the top commonly occurring cancers worldwide, generally showing no early symptoms until the tumor becomes large enough; therefore, the mortality rate is relatively high [18–20]. Thus, it is necessary to investigate the carcinogenesis of KIRC and to identify useful biomarkers for its early diagnosis. However, limited knowledge so far is established about the pathogenesis and carcinogenesis of KIRC. In addition, not many molecular markers for clinical practice have been validated. Advanced high-throughput sequencing and bioinformatics technology make it possible to select effective biomarkers [21]. The RNA sequencing data and clinical annotations of over five hundred KIRC cases are freely available on the TCGA database. Taking advantage of this freely available data from TCGA, we analyzed RNA sequence data for differentially expressed metabolic genes in tumor vs normal tissue samples. Among upregulated and downregulated genes, we identified the top 10 differentially expressed metabolic genes (DEmGs). We believe that metabolic genes have diverse functions in KIRC; still, finding suitable diagnostic and therapeutic markers could be a challenge from the pool of genes with diversified functions.
Previously, studies have estimated immune cell infiltration in the tumor microenvironment of several cancers. A relationship between tumor immune cells and angiogenesis in KIRC samples’ data obtained from TCGA was studied, and RFX2, SOX13, and THRA were identified as the top three MTFs in regulating angiogenesis signature in KIRC patients [4]. Moreover, two independent m6A modification patterns control biological functions, immunological characteristics, and prognoses of KIRC [22]. Autophagy-related protein 5 (ATG5) has been linked with the progression of several cancers including KIRC [23]. In the current analysis, some of the differentially expressed genes including PBRM1, SET2, VHL, and BAP1 showed significant correlation with metabolic pathways in KIRC data. For further deep investigation, we clustered the patients based on the DEmGs; cluster 1 showed a worse overall survival rate as compared with the other clusters; anyhow, the KIRC patients in cluster 3 have advanced tumor stages and have high lymph nodes (higher N1) as compared with those in clusters 1 and 2, showing cancer metastasis and expansion of tumors in cluster 3. It shows a lesser number of metabolic genes in clusters associated with cancer metastasis.
In addition, immune infiltration scores in different clusters show C1 with high scores in stromal and immune classifications. Noteworthy, C1 patients at pathological stages III and IV have high infiltration of T cells along with CD8+ T cells, T follicular helper cells, and macrophages. While also having abundant Tregs. The Tregs have a vital role in immune tolerance and homeostasis [24]. In many cancers such as colon cancer, breast cancer, and pancreas cancer, the increased percentage of Tregs is associated with the poor prognosis of cancer [25, 26]. The M0 macrophage induces the invasion and proliferation of cells [27], and elevated levels of the macrophages are associated with poor prognosis in RCC [28]. Likewise, CD8+ T cells are known as the key antitumor cells and the top choice for targeted immune cell therapy for cancers [29]. Although C1 has the highest infiltration of CD8+ T cells than other clusters, it had the worst overall survival.

Herba Cistanche
We used three methods CIBERSORT, MCP, and ssGSEA to study the immune cell infiltration in the KIRC tumor microenvironment. The traditional method to measure the tumor immune infiltration is through histology on tissue sections and immune subsets inferred by immunohistochemistry of individual markers. However, there are several limitations where immunohistochemistry cannot identify many immune populations and performs poorly at capturing functional phenotypes (e.g., activated vs. resting lymphocytes). Therefore, we utilized CIBERSORT, a computational approach developed by [30] that addresses the challenges faced by immunohistochemistry. Apart from CIBERSORT, we employed other algorithmic packages to check the status of immune infiltration. This is because CIBERSORT measures only intrasample proportions of immune cell populations that can be resolved by another package such as the MCP-counter which can estimate the cells' population in abundance that enables an intersample comparison of infiltrating cells in the tumor microenvironment [31]. To complement CIBERSORT and MCP analyses, we applied ssGSEA to quantify infiltration levels for immune cell types implemented in R package GSVA [32, 33]. The GSA is a rank-based method that computes an overexpression for a gene’s list of interest relative to all other genes in the genome. The CIBERSORT showed better results as compared to the other two methods; thus, further analyses were performed from data obtained from CIBERSORT.
Moreover, we focused on the underlying mechanism of the KIRC progression by differential expression analysis based on the RNA-seq data. Overall, we targeted abnormal differential expression of the common genes among three clusters. Most of the genes were downregulated except NUDT1 in C1; however, its expression was significantly downregulated through the progression from C1 to C3. Thus, NUDT1 was further validated for its role in KIRC progression. The siRNA-mediated inhibition of NUDTI gene expression in two KIRC cell lines (786-O and ACHN) reduced the cell viability and cell migration and increased apoptosis, which confirms its role in tumor progression. Previously, it has been reported that the level of NUDT1 expression correlated with the tumor grade, stage, size, differentiation, degree of vascular invasion, overall survival (OS), and disease-free survival (DFS) in HCC patients, also predicted as a prognostic marker with therapeutic potentials in HCC patients [34]. Overexpressing NUDT1 in pulmonary arterial hypertension reduces oxidative stress and DNA damage, hence promoting cell proliferation and reducing apoptosis [35]. It has been shown that patients with oral squamous cell carcinoma (OSCC) having high expression of NUDT1 have shown a poor survival rate [36]. Based on the fact that not enough literature is available about NUDTI’s role in cancers and, so far, no study has ever reported its role in KIRC, therefore, we are reporting for the first time the role of NUDTI in KIRC progression. The current study had some limitations; although our research found that the signature might be associated with immunotherapy of KIRC, the efficacy of the signature could not be validated due to the lack of data, the potential underlying mechanisms, and functional roles of NUDT1 in KIRC and clinical practice need further exploration.

Cistanche tubulosa
Conclusions
We screened dysregulated metabolic genes between normal and tumor tissues and explored their function. WGCNA analysis identified a group of genes correlated with the survival status of KIRC. Consensus clustering based on survival-related genes demonstrated three clusters with different survival rates and immune infiltration patterns. NUDT1 negatively correlated with survival, and further analyses revealed that the knockdown of NUDT1 inhibits the proliferation and migration of tumor cells. Of note, a prediction model was constructed based on survival-related genes, which showed high efficiency in predicting the survival of KIRC. In conclusion, we performed an exhaustive analysis of metabolic genes in KIRC and identified NUDT1 as an oncogene that could be used as a therapeutic and prognostic target.
The Effect of Cistanche Extract on kidney transparency
There is limited scientific evidence on the effect of Cistanche on kidney transparency. However, some studies suggest that Cistanche extract may have a positive impact on kidney health.
The kidneys are responsible for filtering waste and toxins from the body, and their health is essential for overall well-being. Studies have shown that Cistanche extract has potent antioxidant and anti-inflammatory properties, which can help protect against oxidative stress and inflammation in the kidneys.
In traditional Chinese medicine, Cistanche is also used to tonify the kidneys and improve their function. Some practitioners believe that regular consumption of Cistanche extract can help improve kidney transparency, although this claim is not backed up by robust clinical research.
Therefore, more rigorous research is required to fully understand the potential benefits of Cistanche extract on kidney transparency. Nonetheless, as with any dietary supplement or alternative therapy, it's important to consult with a healthcare professional before incorporating new additions into your diet, particularly if you have pre-existing medical conditions.
References
[18] R. L. Siegel, K. D. Miller, and A. Jemal, “Cancer statistics, 2019,” CA: A Cancer Journal for Clinicians, vol. 69, no. 1, pp. 7–34, 2019.
[19] R. L. Siegel, K. D. Miller, H. E. Fuchs, and A. Jemal, “Cancer statistics, 2021,” CA: A Cancer Journal for Clinicians, vol. 71, no. 1, pp. 7–33, 2021.
[20] J. C. Angulo and O. Shapiro, “The changing therapeutic landscape of metastatic renal cancer,” Cancers, vol. 11, no. 9, p. 1227, 2019.
[21] A. K. Sharma, “Emerging trends in biomarker discovery: ease of prognosis and prediction in cancer,” Seminars in Cancer Biology, vol. 52, Part 1, pp. iii–iv, 2018.
[22] H. Li, J. Hu, A. Yu et al., “RNA modification of N6- methyladenosine predicts immune phenotypes and therapeutic opportunities in kidney renal clear cell carcinoma,” Frontiers in Oncology, vol. 11, article 642159, 2021.
[23] C. Xu, Y. Zang, Y. Zhao, et al., “Comprehensive pan-cancer analysis confirmed that ATG5 promoted the maintenance of tumor metabolism and the occurrence of tumor immune escape,” Frontiers in Oncology, vol. 11, article 652211, 2021.
[24] Y. Takeuchi and H. Nishikawa, “Roles of regulatory T cells in cancer immunity,” International Immunology, vol. 28, no. 8, pp. 401–409, 2016.
[25] C. Zhuo, Y. Xu, M. Ying, et al., “FOXP3+ Tregs: heterogeneous phenotypes and conflicting impacts on survival outcomes in patients with colorectal cancer,” Immunologic Research, vol. 61, no. 3, pp. 338–347, 2015.
[26] H. Wang, F. Franco, and P. C. Ho, “Metabolic regulation of Tregs in cancer: opportunities for immunotherapy,” Trends Cancer, vol. 3, no. 8, pp. 583–592, 2017.
[27] B. Z. Qian and J. W. Pollard, “Macrophage diversity enhances tumor progression and metastasis,” Cell, vol. 141, no. 1, pp. 39–51, 2010.
[28] A. Hajiran, N. Chakiryan, A. M. Aydin, et al., “Reconnaissance of tumor immune microenvironment spatial heterogeneity in metastatic renal cell carcinoma and correlation with immunotherapy response,” Clinical and Experimental Immunology, vol. 204, no. 1, pp. 96–106, 2021.
[29] B. Farhood, M. Najafi, and K. Mortezaee, “CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: a review,” Journal of Cellular Physiology, vol. 234, no. 6, pp. 8509–8521, 2019.
[30] A. M. Newman, C. L. Liu, M. R. Green, et al., “Robust enumeration of cell subsets from tissue expression profiles,” Nature Methods, vol. 12, no. 5, pp. 453–457, 2015.
[31] E. Becht, N. A. Giraldo, L. Lacroix, et al., “Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression,” Genome Biology, vol. 17, no. 1, p. 218, 2016.
[32] D. A. Barbie, P. Tamayo, J. S. Boehm, et al., “Systematic RNA interference reveals that oncogenic KRAS -driven cancers require TBK1,” Nature, vol. 462, no. 7269, pp. 108–112, 2009.
[33] S. Hanzelmann, R. Castelo, and J. Guinney, “GSVA: gene set variation analysis for microarray and RNA-seq data,” BMC Bioinformatics, vol. 14, no. 1, 2013.
[34] Q. Ou, N. Ma, Z. Yu et al., “Nudix hydrolase 1 is a prognostic biomarker in hepatocellular carcinoma,” Aging, vol. 12, no. 8, pp. 7363–7379, 2020.
[35] G. Vitry, R. Paulin, Y. Grobs, et al., “Oxidized DNA precursors cleanup by NUDT1 contributes to vascular remodeling in pulmonary arterial hypertension,” American Journal of Respiratory and Critical Care Medicine, vol. 203, no. 5, pp. 614–627, 2021.
[36] Y. Shen, L. Zhang, S. Piao, et al., “NUDT1: a potential independent predictor for the prognosis of patients with oral squamous cell carcinoma,” Journal of Oral Pathology & Medicine, vol. 49, no. 3, pp. 210–218, 2020.
Junwei Xie,1,2,3,4,5 Lingang Cui,6 Shaokang Pan,1,2,3,4,5 Dongwei Liu,1,2,3,4,5 Fengxun Liu,1,2,3,4,5 and Zhangsuo Liu 1,2,3,4,5
1 Department of Nephrology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, China
2 Research Institute of Nephrology, Zhengzhou University, Zhengzhou 450052, China
3 Research Center for Kidney Disease, Zhengzhou, 450052 Henan, China
4 Key Laboratory of Precision Diagnosis and Treatment for Chronic Kidney Disease in Henan Province, Zhengzhou 450052, China
5 Core Unit of National Clinical Medical Research Center of Kidney Disease, Zhengzhou 450052, China
6 Department of Urology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, China






